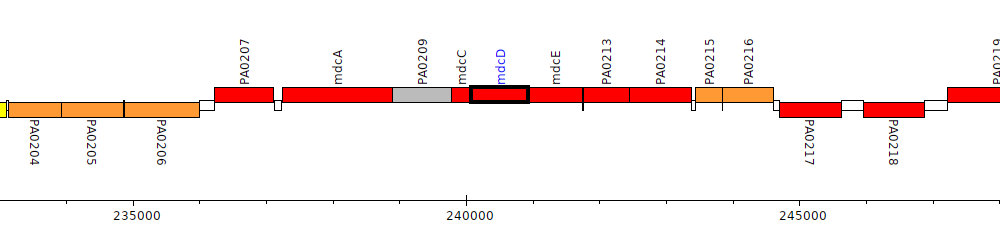
Pseudomonas aeruginosa PAO1, PA0211 (mdcD)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0044248 | cellular catabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0015976 | carbon utilization | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0005975 | carbohydrate metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03133
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016831 | carboxy-lyase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03133
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF52096 | ClpP/crotonase | IPR029045 | ClpP/crotonase-like domain superfamily | 4 | 236 | 8.71E-37 |
| PANTHER | PTHR42995 | ACETYL-COENZYME A CARBOXYLASE CARBOXYL TRANSFERASE SUBUNIT BETA, CHLOROPLASTIC | - | - | 18 | 214 | 3.6E-21 |
| Pfam | PF01039 | Carboxyl transferase domain | IPR034733 | Acetyl-CoA carboxylase | 18 | 235 | 1.6E-14 |
| NCBIfam | TIGR03133 | JCVI: biotin-independent malonate decarboxylase subunit beta | IPR017556 | Biotin-independent malonate decarboxylase, beta subunit | 11 | 281 | 1.1E-116 |
| Gene3D | G3DSA:3.90.226.10 | - | - | - | 3 | 240 | 4.1E-47 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.