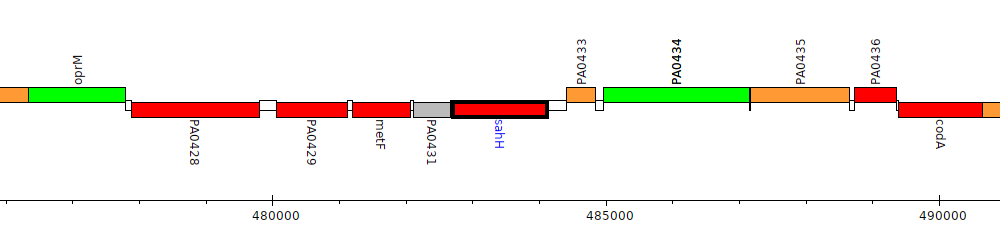
Pseudomonas aeruginosa PAO1, PA0432 (sahH)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae00270 | Cysteine and methionine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| NCBIfam | TIGR00936 | JCVI: adenosylhomocysteinase | IPR000043 | Adenosylhomocysteinase-like | 13 | 461 | 0.0 |
| Pfam | PF05221 | S-adenosyl-L-homocysteine hydrolase | IPR000043 | Adenosylhomocysteinase-like | 12 | 149 | 4.8E-72 |
| FunFam | G3DSA:3.40.50.1480:FF:000006 | Adenosylhomocysteinase | - | - | 2 | 150 | 4.8E-88 |
| FunFam | G3DSA:3.40.50.720:FF:000155 | Adenosylhomocysteinase | - | - | 206 | 377 | 6.3E-115 |
| Hamap | MF_00563 | S-inosyl-L-homocysteine hydrolase [ahcY]. | IPR000043 | Adenosylhomocysteinase-like | 8 | 461 | 43.834774 |
| SMART | SM00996 | AdoHcyase_2 | IPR000043 | Adenosylhomocysteinase-like | 12 | 468 | 0.0 |
| CDD | cd00401 | SAHH | IPR000043 | Adenosylhomocysteinase-like | 22 | 456 | 0.0 |
| Pfam | PF00670 | S-adenosyl-L-homocysteine hydrolase, NAD binding domain | IPR015878 | S-adenosyl-L-homocysteine hydrolase, NAD binding domain | 199 | 381 | 1.0E-55 |
| SMART | SM00997 | AdoHcyase_NAD_2 | IPR015878 | S-adenosyl-L-homocysteine hydrolase, NAD binding domain | 199 | 381 | 4.8E-82 |
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 206 | 376 | 2.2E-76 |
| FunFam | G3DSA:3.40.50.1480:FF:000007 | Adenosylhomocysteinase | - | - | 151 | 205 | 2.9E-37 |
| Gene3D | G3DSA:3.40.50.1480 | - | IPR042172 | Adenosylhomocysteinase-like superfamily | 377 | 469 | 5.2E-31 |
| Gene3D | G3DSA:3.40.50.1480 | - | IPR042172 | Adenosylhomocysteinase-like superfamily | 2 | 150 | 1.4E-71 |
| PANTHER | PTHR23420 | ADENOSYLHOMOCYSTEINASE | IPR000043 | Adenosylhomocysteinase-like | 9 | 469 | 0.0 |
| FunFam | G3DSA:3.40.50.1480:FF:000013 | Adenosylhomocysteinase | - | - | 377 | 469 | 4.5E-61 |
| SUPERFAMILY | SSF51735 | NAD(P)-binding Rossmann-fold domains | IPR036291 | NAD(P)-binding domain superfamily | 199 | 381 | 1.47E-50 |
| Gene3D | G3DSA:3.40.50.1480 | - | IPR042172 | Adenosylhomocysteinase-like superfamily | 151 | 205 | 1.7E-22 |
| SUPERFAMILY | SSF52283 | Formate/glycerate dehydrogenase catalytic domain-like | - | - | 10 | 469 | 8.5E-108 |
| PIRSF | PIRSF001109 | SAHH | IPR000043 | Adenosylhomocysteinase-like | 2 | 469 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.