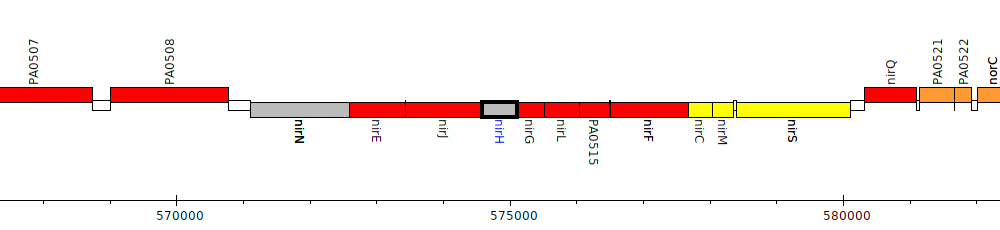
Pseudomonas aeruginosa PAO1, PA0512 (nirH)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0009108 | obsolete coenzyme biosynthetic process | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
8982003 | Reviewed by curator |
| Biological Process | GO:0006783 | heme biosynthetic process | Inferred from Mutant Phenotype | ECO:0000016 loss-of-function mutant phenotype evidence |
8982003 | Reviewed by curator |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Denitrification |
ECO:0000037
not_recorded |
|||
| PseudoCAP | Biosynthesis of heme d1 |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF17805 | AsnC-like ligand binding domain | IPR040523 | Siroheme decarboxylase , AsnC-like ligand binding domain | 69 | 157 | 2.7E-22 |
| Gene3D | G3DSA:3.30.70.3460 | - | - | - | 27 | 170 | 2.0E-43 |
| FunFam | G3DSA:3.30.70.3460:FF:000002 | Heme d1 biosynthesis protein NirH | - | - | 27 | 170 | 8.8E-95 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.