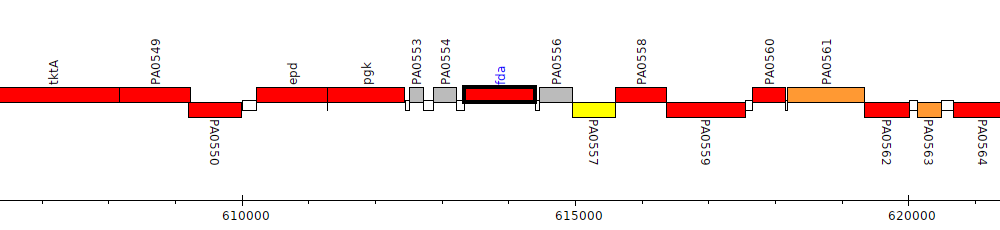
Pseudomonas aeruginosa PAO1, PA0555 (fda)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004332 | fructose-bisphosphate aldolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01521
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0005975 | carbohydrate metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF001359
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006096 | glycolytic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01521
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016832 | aldehyde-lyase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF001359
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008270 | zinc ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF001359
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | PWY-5484 | glycolysis II (from fructose 6-phosphate) | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01200 | Carbon metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00680 | Methane metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | GLYCOLYSIS | glycolysis I (from glucose 6-phosphate) | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | GLYCOLYSIS+CITRIC-ACID-PWY | glycolysis + citric acid pathway | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae00051 | Fructose and mannose metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00010 | Glycolysis / Gluconeogenesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | ANAGLYCOLYSIS-PWY | glycolysis III (from glucose) | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | GLYCOLYSIS-E-D | superpathway of glycolysis and Entner-Doudoroff | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCAP | Pentose phosphate cycle |
ECO:0000037
not_recorded |
|||
| PseudoCAP | Fructose and mannose metabolism |
ECO:0000037
not_recorded |
|||
| KEGG | pae01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Carbon fixation |
ECO:0000037
not_recorded |
|||
| PseudoCyc | GLYCOLYSIS-TCA-GLYOX-BYPASS | superpathway of glycolysis, pyruvate dehydrogenase, TCA, and glyoxylate bypass | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | P461-PWY | hexitol fermentation to lactate, formate, ethanol and acetate | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | GLUCONEO-PWY | gluconeogenesis I | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Glycolysis / Gluconeogenesis |
ECO:0000037
not_recorded |
|||
| KEGG | pae00030 | Pentose phosphate pathway | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01230 | Biosynthesis of amino acids | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF01116 | Fructose-bisphosphate aldolase class-II | IPR000771 | Fructose-bisphosphate aldolase, class-II | 5 | 327 | 7.3E-92 |
| FunFam | G3DSA:3.20.20.70:FF:000111 | Fructose-1,6-bisphosphate aldolase | - | - | 1 | 345 | 0.0 |
| NCBIfam | TIGR00167 | JCVI: ketose-bisphosphate aldolase | IPR000771 | Fructose-bisphosphate aldolase, class-II | 1 | 329 | 2.8E-97 |
| CDD | cd00947 | TBP_aldolase_IIB | IPR000771 | Fructose-bisphosphate aldolase, class-II | 6 | 327 | 0.0 |
| PANTHER | PTHR30304 | D-TAGATOSE-1,6-BISPHOSPHATE ALDOLASE | - | - | 2 | 331 | 1.7E-122 |
| PIRSF | PIRSF001359 | F_bP_aldolase_II | IPR000771 | Fructose-bisphosphate aldolase, class-II | 1 | 329 | 1.1E-83 |
| NCBIfam | TIGR01521 | JCVI: fructose-bisphosphate aldolase class II | IPR006412 | Fructose-bisphosphate aldolase, class II, Calvin cycle subtype | 3 | 349 | 0.0 |
| Gene3D | G3DSA:3.20.20.70 | Aldolase class I | IPR013785 | Aldolase-type TIM barrel | 1 | 354 | 0.0 |
| SUPERFAMILY | SSF51569 | Aldolase | - | - | 3 | 328 | 7.77E-107 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.