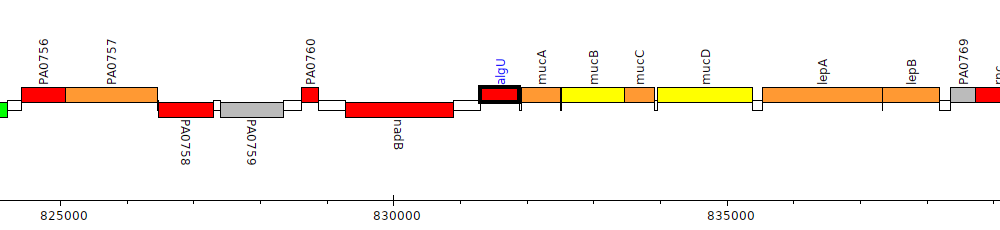
Pseudomonas aeruginosa PAO1, PA0762 (algU)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:1900189 | positive regulation of cell adhesion involved in single-species biofilm formation | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
20348252 | Reviewed by curator |
| Biological Process | GO:0071236 | cellular response to antibiotic | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
18838593 | Reviewed by curator |
| Biological Process | GO:0036460 | cellular response to cell envelope stress | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
19226327 | Reviewed by curator |
| Biological Process | GO:1902201 | negative regulation of bacterial-type flagellum-dependent cell motility | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
16291668 | Reviewed by curator |
| Molecular Function | GO:0016987 | sigma factor activity | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
7644517 | Reviewed by curator |
| Biological Process | GO:0032885 | regulation of polysaccharide biosynthetic process | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
14645293 | Reviewed by curator |
| Biological Process | GO:2000154 | obsolete negative regulation of flagellar cell motility | |
|||
| Biological Process | GO:1900036 | positive regulation of cellular response to heat | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
9159526 | Reviewed by curator |
| Biological Process | GO:1900233 | positive regulation of single-species biofilm formation on inanimate substrate | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
20348252 | Reviewed by curator |
| Biological Process | GO:1902884 | positive regulation of response to oxidative stress | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
7961422 | Reviewed by curator |
| Biological Process | GO:0032885 | regulation of polysaccharide biosynthetic process | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
7928961 | Reviewed by curator |
| Biological Process | GO:0009405 | pathogenesis | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
8698507 | Reviewed by curator |
| Cellular Component | GO:0000345 | cytosolic DNA-directed RNA polymerase complex | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
7644517 | Reviewed by curator |
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02939
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016987 | sigma factor activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02939
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006352 | DNA-templated transcription, initiation |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02939
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003700 | DNA-binding transcription factor activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02939
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02939
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Alginate biosynthesis |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| FunFam | G3DSA:1.10.10.10:FF:000043 | RNA polymerase sigma factor | - | - | 122 | 192 | 1.0E-39 |
| Gene3D | G3DSA:1.10.1740.10 | - | - | - | 1 | 121 | 4.4E-36 |
| PANTHER | PTHR43133 | RNA POLYMERASE ECF-TYPE SIGMA FACTO | IPR039425 | RNA polymerase sigma-70 like | 3 | 189 | 2.8E-47 |
| SUPERFAMILY | SSF88946 | Sigma2 domain of RNA polymerase sigma factors | IPR013325 | RNA polymerase sigma factor, region 2 | 1 | 108 | 1.38E-32 |
| SUPERFAMILY | SSF88659 | Sigma3 and sigma4 domains of RNA polymerase sigma factors | IPR013324 | RNA polymerase sigma factor, region 3/4-like | 123 | 187 | 7.85E-18 |
| NCBIfam | TIGR02937 | JCVI: sigma-70 family RNA polymerase sigma factor | IPR014284 | RNA polymerase sigma-70 like domain | 21 | 186 | 2.0E-30 |
| Gene3D | G3DSA:1.10.10.10 | - | IPR036388 | Winged helix-like DNA-binding domain superfamily | 122 | 192 | 2.2E-23 |
| Pfam | PF08281 | Sigma-70, region 4 | IPR013249 | RNA polymerase sigma factor 70, region 4 type 2 | 132 | 181 | 3.1E-18 |
| NCBIfam | TIGR02939 | JCVI: RNA polymerase sigma factor RpoE | IPR014286 | RNA polymerase sigma-70 RpoE type | 3 | 190 | 2.4E-98 |
| CDD | cd06171 | Sigma70_r4 | - | - | 132 | 182 | 6.40095E-13 |
| FunFam | G3DSA:1.10.1740.10:FF:000001 | RNA polymerase sigma factor | - | - | 1 | 121 | 1.2E-65 |
| Pfam | PF04542 | Sigma-70 region 2 | IPR007627 | RNA polymerase sigma-70 region 2 | 25 | 92 | 1.1E-17 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.