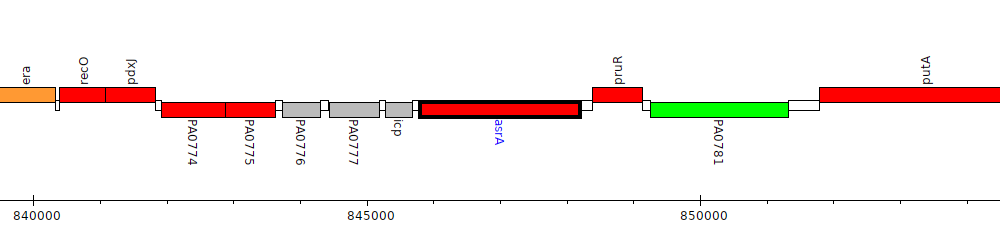
Pseudomonas aeruginosa PAO1, PA0779 (asrA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0071236 | cellular response to antibiotic | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
21357290 | Reviewed by curator |
| Biological Process | GO:0080164 | regulation of nitric oxide metabolic process | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
18043907 | Reviewed by curator |
| Molecular Function | GO:0004176 | ATP-dependent peptidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF05362
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006508 | proteolysis |
Inferred from Sequence Model
Term mapped from: InterPro:PF05362
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00004
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0043565 | sequence-specific DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01973
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004252 | serine-type endopeptidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF05362
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016887 | ATPase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00004
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0030163 | protein catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR43718
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF05362 | Lon protease (S16) C-terminal proteolytic domain | IPR008269 | Peptidase S16, Lon proteolytic domain | 597 | 797 | 2.8E-79 |
| PANTHER | PTHR43718 | LON PROTEASE | IPR027065 | Lon protease | 31 | 594 | 0.0 |
| Gene3D | G3DSA:1.20.5.5270 | - | - | - | 279 | 328 | 3.7E-15 |
| Gene3D | G3DSA:3.30.230.10 | - | IPR014721 | Small ribosomal subunit protein uS5 domain 2-type fold, subgroup | 615 | 799 | 3.7E-66 |
| PRINTS | PR00830 | Endopeptidase La (Lon) serine protease (S16) signature | - | - | 729 | 748 | 1.6E-47 |
| Hamap | MF_01973 | Lon protease [lon]. | IPR027543 | Lon protease, bacterial | 32 | 798 | 59.922005 |
| FunFam | G3DSA:1.20.58.1480:FF:000008 | Lon protease | - | - | 146 | 270 | 1.0E-72 |
| SMART | SM00464 | lon_5 | IPR003111 | Lon protease, N-terminal domain | 31 | 228 | 8.5E-23 |
| SUPERFAMILY | SSF88697 | PUA domain-like | IPR015947 | PUA-like superfamily | 32 | 227 | 6.46E-19 |
| Coils | Coil | Coil | - | - | 225 | 245 | - |
| Pfam | PF02190 | ATP-dependent protease La (LON) substrate-binding domain | IPR003111 | Lon protease, N-terminal domain | 32 | 227 | 1.4E-21 |
| PIRSF | PIRSF001174 | Lon_proteas | IPR004815 | Lon protease, bacterial/eukaryotic-type | 21 | 799 | 0.0 |
| Gene3D | G3DSA:3.40.50.300 | - | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 340 | 518 | 1.0E-61 |
| FunFam | G3DSA:3.40.50.300:FF:000021 | Lon protease homolog | - | - | 340 | 518 | 7.9E-91 |
| SUPERFAMILY | SSF52540 | P-loop containing nucleoside triphosphate hydrolases | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 337 | 614 | 1.01E-48 |
| PRINTS | PR00830 | Endopeptidase La (Lon) serine protease (S16) signature | - | - | 752 | 770 | 1.6E-47 |
| Gene3D | G3DSA:1.20.58.1480 | - | - | - | 146 | 270 | 1.6E-31 |
| NCBIfam | TIGR00763 | JCVI: endopeptidase La | IPR004815 | Lon protease, bacterial/eukaryotic-type | 34 | 797 | 0.0 |
| CDD | cd19500 | RecA-like_Lon | - | - | 340 | 521 | 5.8384E-122 |
| Pfam | PF00004 | ATPase family associated with various cellular activities (AAA) | IPR003959 | ATPase, AAA-type, core | 379 | 517 | 1.7E-22 |
| PRINTS | PR00830 | Endopeptidase La (Lon) serine protease (S16) signature | - | - | 383 | 402 | 1.6E-47 |
| Gene3D | G3DSA:1.10.8.60 | - | - | - | 519 | 611 | 1.3E-27 |
| FunFam | G3DSA:1.20.5.5270:FF:000002 | Lon protease homolog | - | - | 279 | 328 | 5.7E-17 |
| Gene3D | G3DSA:2.30.130.40 | - | IPR046336 | Lon protease, N-terminal domain superfamily | 25 | 139 | 1.3E-13 |
| PRINTS | PR00830 | Endopeptidase La (Lon) serine protease (S16) signature | - | - | 699 | 718 | 1.6E-47 |
| SUPERFAMILY | SSF54211 | Ribosomal protein S5 domain 2-like | IPR020568 | Ribosomal protein uS5 domain 2-type superfamily | 620 | 797 | 9.48E-47 |
| SMART | SM00382 | AAA_5 | IPR003593 | AAA+ ATPase domain | 375 | 520 | 3.1E-12 |
| PRINTS | PR00830 | Endopeptidase La (Lon) serine protease (S16) signature | - | - | 620 | 636 | 1.6E-47 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 1 | 22 | - |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.