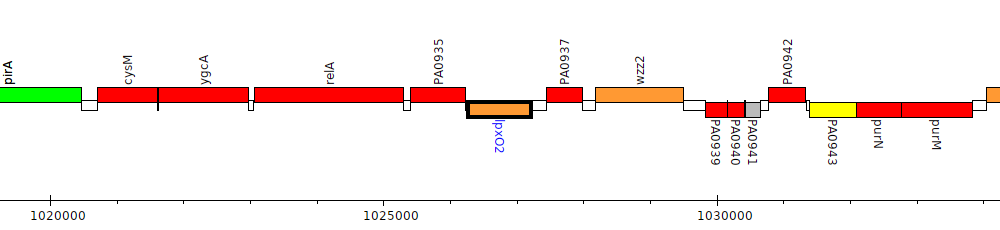
Pseudomonas aeruginosa PAO1, PA0936 (lpxO2)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0009276 | Gram-negative-bacterium-type cell wall | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0003824 | catalytic activity | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0018193 | peptidyl-amino acid modification |
Inferred from Sequence Model
Term mapped from: InterPro:PF05118
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR46332 | ASPARTATE BETA-HYDROXYLASE DOMAIN-CONTAINING PROTEIN 2 | - | - | 18 | 234 | 2.5E-62 |
| SUPERFAMILY | SSF51197 | Clavaminate synthase-like | - | - | 105 | 227 | 3.02E-17 |
| Gene3D | G3DSA:2.60.120.330 | - | IPR027443 | Isopenicillin N synthase-like superfamily | 49 | 235 | 1.0E-46 |
| Pfam | PF05118 | Aspartyl/Asparaginyl beta-hydroxylase | IPR007803 | Aspartyl/asparaginy/proline hydroxylase | 75 | 228 | 1.4E-46 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.