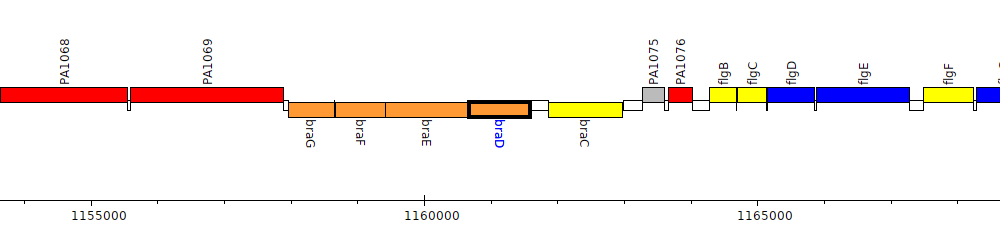
Pseudomonas aeruginosa PAO1, PA1073 (braD)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0016020 | membrane | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Biological Process | GO:0015808 | L-alanine transport | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
18833300 | Reviewed by curator |
| Biological Process | GO:0006810 | transport | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Biological Process | GO:1903806 | L-isoleucine import across plasma membrane | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
18833300 | Reviewed by curator |
| Biological Process | GO:0042941 | D-alanine transport | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
18833300 | Reviewed by curator |
| Molecular Function | GO:0022857 | transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF02653
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PF02653
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055085 | transmembrane transport |
Inferred from Sequence Model
Term mapped from: InterPro:PF02653
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae02010 | ABC transporters | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae02024 | Quorum sensing | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR11795 | BRANCHED-CHAIN AMINO ACID TRANSPORT SYSTEM PERMEASE PROTEIN LIVH | - | - | 1 | 303 | 1.7E-85 |
| Pfam | PF02653 | Branched-chain amino acid transport system / permease component | IPR001851 | ABC transporter, permease | 11 | 291 | 3.4E-54 |
| CDD | cd06582 | TM_PBP1_LivH_like | - | - | 14 | 299 | 6.81417E-66 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.