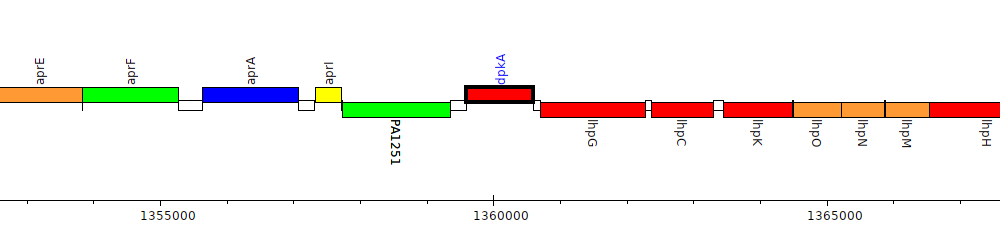
Pseudomonas aeruginosa PAO1, PA1252 (dpkA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0019470 | 4-hydroxyproline catabolic process | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
27145750 | Reviewed by curator |
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF02615
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCyc | P22-PWY | acetyl-CoA assimilation | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | P106-PWY | serine-isocitrate lyase pathway | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | TCA-GLYOX-BYPASS | superpathway of glyoxylate bypass and TCA | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | GLUCONEO-PWY | gluconeogenesis I | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | GLYOXYLATE-BYPASS | glyoxylate cycle | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | ANARESP1-PWY | anaerobic respiration | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | MALATE-ASPARTATE-SHUTTLE-PWY | L-aspartate degradation II | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | GLYCOLYSIS-TCA-GLYOX-BYPASS | superpathway of glycolysis, pyruvate dehydrogenase, TCA, and glyoxylate bypass | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | FERMENTATION-PWY | mixed acid fermentation | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | TCA | TCA cycle I (prokaryotic) | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae00310 | Lysine degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| FunFam | G3DSA:3.30.1370.60:FF:000002 | Malate/L-lactate family dehydrogenase | - | - | 68 | 303 | 2.3E-112 |
| Gene3D | G3DSA:3.30.1370.60 | Hypothetical oxidoreductase yiak; domain 2 | IPR043143 | Malate/L-sulfolactate/L-lactate dehydrogenase-like, NADPH binding domain | 68 | 303 | 7.5E-88 |
| Gene3D | G3DSA:1.10.1530.10 | - | IPR043144 | Malate/L-sulfolactate/L-lactate dehydrogenase-like, alpha-helical domain | 13 | 332 | 7.5E-88 |
| Pfam | PF02615 | Malate/L-lactate dehydrogenase | IPR003767 | Malate/L-lactate dehydrogenase-like | 5 | 329 | 4.2E-101 |
| SUPERFAMILY | SSF89733 | L-sulfolactate dehydrogenase-like | IPR036111 | Malate/L-sulfolactate/L-lactate dehydrogenase-like superfamily | 1 | 333 | 2.35E-101 |
| PANTHER | PTHR11091 | OXIDOREDUCTASE-RELATED | IPR003767 | Malate/L-lactate dehydrogenase-like | 3 | 332 | 4.9E-74 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.