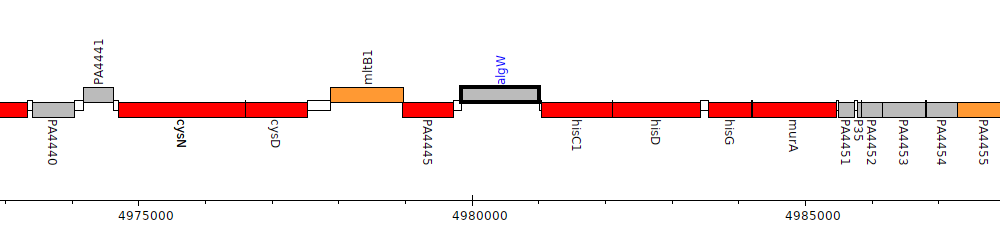
Pseudomonas aeruginosa PAO1, PA4446 (algW)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0008233 | peptidase activity | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
19298369 | Reviewed by curator |
| Biological Process | GO:0050896 | response to stimulus | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Biological Process | GO:0044267 | cellular protein metabolic process | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Molecular Function | GO:0004252 | serine-type endopeptidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR00834
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005515 | protein binding |
Inferred from Sequence Model
Term mapped from: InterPro:SM00228
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006508 | proteolysis |
Inferred from Sequence Model
Term mapped from: InterPro:PR00834
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM00228 | pdz_new | IPR001478 | PDZ domain | 280 | 362 | 1.3E-12 |
| Gene3D | G3DSA:2.30.42.10 | - | IPR036034 | PDZ superfamily | 278 | 376 | 2.9E-25 |
| PRINTS | PR00834 | HtrA/DegQ protease family signature | IPR001940 | Peptidase S1C | 177 | 201 | 5.6E-54 |
| Gene3D | G3DSA:2.40.10.120 | - | - | - | 53 | 274 | 4.7E-77 |
| Pfam | PF13180 | PDZ domain | IPR001478 | PDZ domain | 281 | 370 | 6.7E-15 |
| CDD | cd00987 | PDZ_serine_protease | - | - | 280 | 368 | 3.53221E-24 |
| PANTHER | PTHR22939 | SERINE PROTEASE FAMILY S1C HTRA-RELATED | - | - | 32 | 369 | 1.9E-73 |
| FunFam | G3DSA:2.40.10.10:FF:000001 | Periplasmic serine protease DegS | - | - | 168 | 279 | 4.8E-41 |
| PRINTS | PR00834 | HtrA/DegQ protease family signature | IPR001940 | Peptidase S1C | 212 | 229 | 5.6E-54 |
| Pfam | PF13365 | Trypsin-like peptidase domain | - | - | 107 | 241 | 1.1E-29 |
| Pfam | PF02163 | Peptidase family M50 | IPR008915 | Peptidase M50 | 262 | 370 | 8.8E-8 |
| PRINTS | PR00834 | HtrA/DegQ protease family signature | IPR001940 | Peptidase S1C | 320 | 332 | 5.6E-54 |
| PRINTS | PR00834 | HtrA/DegQ protease family signature | IPR001940 | Peptidase S1C | 137 | 157 | 5.6E-54 |
| SUPERFAMILY | SSF50156 | PDZ domain-like | IPR036034 | PDZ superfamily | 238 | 363 | 1.71E-22 |
| PRINTS | PR00834 | HtrA/DegQ protease family signature | IPR001940 | Peptidase S1C | 234 | 251 | 5.6E-54 |
| SUPERFAMILY | SSF50494 | Trypsin-like serine proteases | IPR009003 | Peptidase S1, PA clan | 51 | 272 | 6.4E-63 |
| PRINTS | PR00834 | HtrA/DegQ protease family signature | IPR001940 | Peptidase S1C | 116 | 128 | 5.6E-54 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.