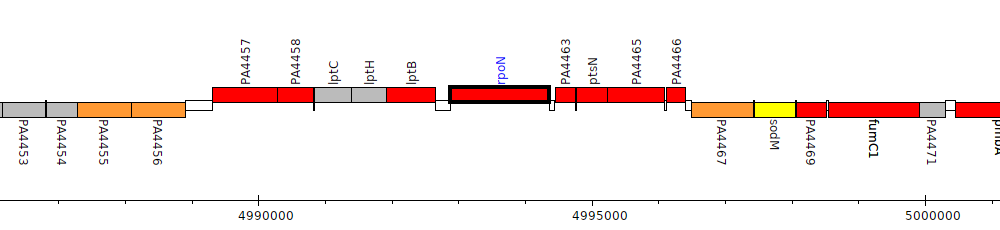
Pseudomonas aeruginosa PAO1, PA4462 (rpoN)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:1900191 | negative regulation of single-species biofilm formation | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
15231779 | Reviewed by curator |
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Biological Process | GO:0006352 | DNA-templated transcription, initiation |
Inferred from Sequence Model
Term mapped from: InterPro:PF04963
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF04963
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016987 | sigma factor activity |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR32248
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0001216 | DNA-binding transcription activator activity |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR32248
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae02020 | Two-component system | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| NCBIfam | TIGR02395 | JCVI: RNA polymerase factor sigma-54 | IPR000394 | RNA polymerase sigma factor 54 | 10 | 494 | 0.0 |
| PRINTS | PR00045 | Sigma-54 factor signature | IPR000394 | RNA polymerase sigma factor 54 | 396 | 408 | 1.4E-32 |
| FunFam | G3DSA:1.10.10.60:FF:000045 | RNA polymerase sigma-54 factor | - | - | 430 | 494 | 6.8E-38 |
| PIRSF | PIRSF000774 | RpoN | IPR000394 | RNA polymerase sigma factor 54 | 1 | 497 | 0.0 |
| Pfam | PF04963 | Sigma-54 factor, core binding domain | IPR007046 | RNA polymerase sigma factor 54, core-binding domain | 129 | 323 | 5.2E-63 |
| PRINTS | PR00045 | Sigma-54 factor signature | IPR000394 | RNA polymerase sigma factor 54 | 470 | 488 | 1.4E-32 |
| Gene3D | G3DSA:1.10.10.60 | - | - | - | 430 | 494 | 2.8E-25 |
| FunFam | G3DSA:1.10.10.1330:FF:000001 | RNA polymerase sigma-54 factor | - | - | 140 | 213 | 7.5E-36 |
| PANTHER | PTHR32248 | RNA POLYMERASE SIGMA-54 FACTOR | IPR000394 | RNA polymerase sigma factor 54 | 1 | 496 | 0.0 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 42 | 100 | - |
| Pfam | PF00309 | Sigma-54 factor, Activator interacting domain (AID) | IPR000394 | RNA polymerase sigma factor 54 | 5 | 48 | 9.1E-17 |
| Gene3D | G3DSA:1.10.10.1330 | - | IPR038709 | RNA polymerase sigma-54 factor, core-binding domain superfamily | 140 | 213 | 2.6E-23 |
| Coils | Coil | Coil | - | - | 252 | 272 | - |
| Pfam | PF04552 | Sigma-54, DNA binding domain | IPR007634 | RNA polymerase sigma factor 54, DNA-binding | 337 | 495 | 3.0E-70 |
| PRINTS | PR00045 | Sigma-54 factor signature | IPR000394 | RNA polymerase sigma factor 54 | 206 | 223 | 1.4E-32 |
| PRINTS | PR00045 | Sigma-54 factor signature | IPR000394 | RNA polymerase sigma factor 54 | 412 | 423 | 1.4E-32 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 72 | 89 | - |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.