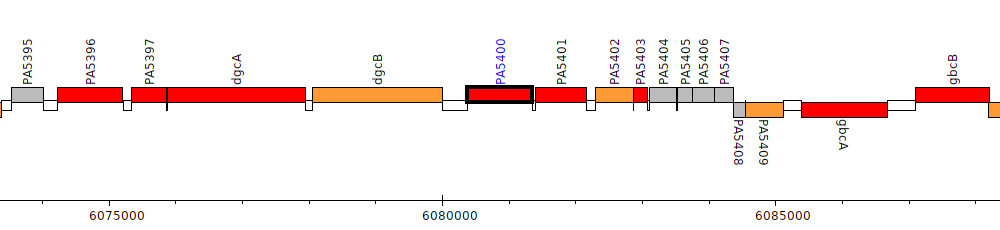
Pseudomonas aeruginosa PAO1, PA5400
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0009055 | electron transfer activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000089
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0050660 | flavin adenine dinucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000089
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM00893 | ETF_2 | IPR014730 | Electron transfer flavoprotein, alpha/beta-subunit, N-terminal | 13 | 187 | 1.2E-6 |
| PIRSF | PIRSF000089 | Electra_flavoP_a | IPR001308 | Electron transfer flavoprotein alpha subunit/FixB | 10 | 321 | 2.2E-69 |
| Gene3D | G3DSA:3.40.50.620 | HUPs | IPR014729 | Rossmann-like alpha/beta/alpha sandwich fold | 12 | 177 | 9.2E-16 |
| SUPERFAMILY | SSF52402 | Adenine nucleotide alpha hydrolases-like | - | - | 13 | 137 | 3.77E-20 |
| PANTHER | PTHR43153 | ELECTRON TRANSFER FLAVOPROTEIN ALPHA | IPR001308 | Electron transfer flavoprotein alpha subunit/FixB | 13 | 314 | 9.9E-43 |
| Pfam | PF01012 | Electron transfer flavoprotein domain | IPR014730 | Electron transfer flavoprotein, alpha/beta-subunit, N-terminal | 19 | 131 | 8.8E-14 |
| Pfam | PF00766 | Electron transfer flavoprotein FAD-binding domain | IPR014731 | Electron transfer flavoprotein, alpha subunit, C-terminal | 201 | 276 | 3.9E-24 |
| Gene3D | G3DSA:3.40.50.1220 | - | - | - | 194 | 326 | 5.9E-36 |
| SUPERFAMILY | SSF52467 | DHS-like NAD/FAD-binding domain | IPR029035 | DHS-like NAD/FAD-binding domain superfamily | 199 | 314 | 1.13E-33 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.