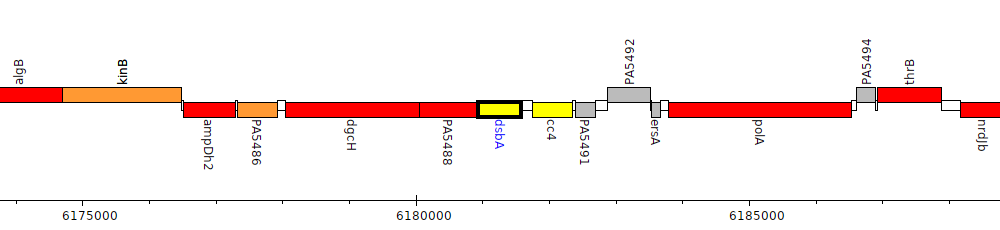
Pseudomonas aeruginosa PAO1, PA5489 (dsbA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006457 | protein folding | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0044267 | cellular protein metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01323
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01503 | Cationic antimicrobial peptide (CAMP) resistance | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF52833 | Thioredoxin-like | IPR036249 | Thioredoxin-like superfamily | 25 | 203 | 4.03E-45 |
| PIRSF | PIRSF001488 | Tdi_protein | IPR023205 | Thiol:disulphide interchange protein DsbA/DsbL | 1 | 209 | 9.8E-78 |
| PANTHER | PTHR35891 | THIOL:DISULFIDE INTERCHANGE PROTEIN DSBA | - | - | 1 | 210 | 1.1E-66 |
| Gene3D | G3DSA:3.40.30.10 | Glutaredoxin | - | - | 20 | 211 | 4.5E-59 |
| CDD | cd03019 | DsbA_DsbA | IPR023205 | Thiol:disulphide interchange protein DsbA/DsbL | 27 | 207 | 1.51295E-72 |
| Pfam | PF01323 | DSBA-like thioredoxin domain | IPR001853 | DSBA-like thioredoxin domain | 96 | 182 | 4.3E-15 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.