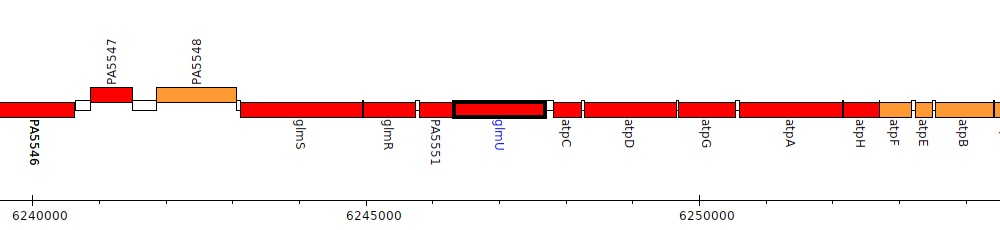
Pseudomonas aeruginosa PAO1, PA5552 (glmU)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003977 | UDP-N-acetylglucosamine diphosphorylase activity | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
8555230 | Reviewed by curator |
| Biological Process | GO:0006048 | UDP-N-acetylglucosamine biosynthetic process | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
8407787 | Reviewed by curator |
| Molecular Function | GO:0019134 | glucosamine-1-phosphate N-acetyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01173
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0000902 | cell morphogenesis |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01173
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003977 | UDP-N-acetylglucosamine diphosphorylase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01173
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005737 | cytoplasm |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01173
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009252 | peptidoglycan biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01173
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006048 | UDP-N-acetylglucosamine biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01173
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000287 | magnesium ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01173
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae00520 | Amino sugar and nucleotide sugar metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | OANTIGEN-PWY | O-antigen building blocks biosynthesis (E. coli) | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Aminosugars metabolism |
ECO:0000037
not_recorded |
|||
| PseudoCyc | KDO-PEP-LIPASYN-PWY | KDO<SUB>2</SUB>-lipid A and peptidoglycan biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | PEP-LIPA-SYN-PWY | peptidoglycan and lipid A precursor biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | UDPNAGSYN-PWY | UDP-N-acetyl-D-glucosamine biosynthesis I | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF14602 | Hexapeptide repeat of succinyl-transferase | IPR001451 | Hexapeptide repeat | 394 | 425 | 2.7E-5 |
| Gene3D | G3DSA:3.90.550.10 | Spore Coat Polysaccharide Biosynthesis Protein SpsA; Chain A | IPR029044 | Nucleotide-diphospho-sugar transferases | 1 | 224 | 5.2E-71 |
| CDD | cd03353 | LbH_GlmU_C | IPR038009 | GlmU, C-terminal LbH domain | 248 | 440 | 1.29457E-121 |
| PANTHER | PTHR43584 | NUCLEOTIDYL TRANSFERASE | - | - | 1 | 420 | 0.0 |
| Pfam | PF12804 | MobA-like NTP transferase domain | IPR025877 | MobA-like NTP transferase | 6 | 121 | 1.8E-24 |
| Hamap | MF_01631 | Bifunctional protein GlmU [glmU]. | IPR005882 | Bifunctional protein GlmU | 2 | 449 | 43.32618 |
| CDD | cd02540 | GT2_GlmU_N_bac | - | - | 5 | 228 | 1.82084E-124 |
| SUPERFAMILY | SSF51161 | Trimeric LpxA-like enzymes | IPR011004 | Trimeric LpxA-like superfamily | 252 | 439 | 2.45E-49 |
| SUPERFAMILY | SSF53448 | Nucleotide-diphospho-sugar transferases | IPR029044 | Nucleotide-diphospho-sugar transferases | 1 | 249 | 1.07E-63 |
| NCBIfam | TIGR01173 | JCVI: UDP-N-acetylglucosamine diphosphorylase/glucosamine-1-phosphate N-acetyltransferase | IPR005882 | Bifunctional protein GlmU | 4 | 451 | 0.0 |
| Pfam | PF00132 | Bacterial transferase hexapeptide (six repeats) | IPR001451 | Hexapeptide repeat | 264 | 296 | 1.3E-7 |
| Gene3D | G3DSA:2.160.10.10 | Hexapeptide repeat proteins | - | - | 225 | 453 | 1.5E-72 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.