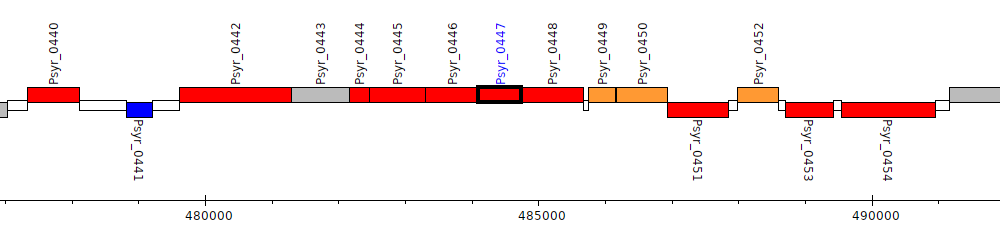
Pseudomonas syringae pv. syringae B728a, Psyr_0447
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016779 | nucleotidyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00650
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | malonate decarboxylase activation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| NCBIfam | TIGR03135 | JCVI: malonate decarboxylase holo-[acyl-carrier-protein] synthase | IPR017557 | Phosphoribosyl-dephospho-CoA transferase | 11 | 204 | 7.7E-53 |
| Hamap | MF_00650 | Phosphoribosyl-dephospho-CoA transferase [mdcG]. | IPR017557 | Phosphoribosyl-dephospho-CoA transferase | 8 | 210 | 34.618305 |
| Pfam | PF10620 | Phosphoribosyl-dephospho-CoA transferase MdcG | IPR017557 | Phosphoribosyl-dephospho-CoA transferase | 11 | 205 | 5.8E-53 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.