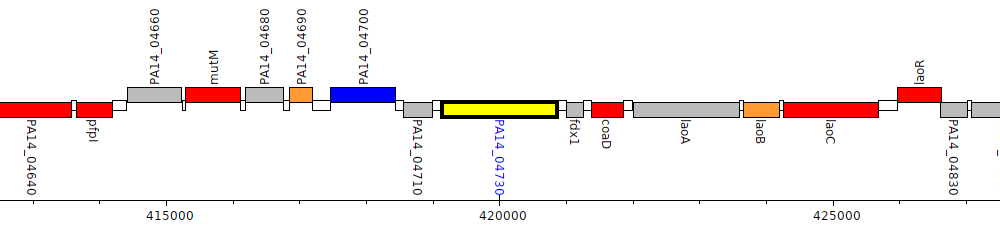
Pseudomonas aeruginosa UCBPP-PA14, PA14_04730
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0036374 | glutathione hydrolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00066
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006751 | glutathione catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00066
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | glutathione-mediated detoxification I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | pau00480 | Glutathione metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau00430 | Taurine and hypotaurine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | γ-glutamyl cycle | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | glutathione-mediated detoxification II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | pau00460 | Cyanoamino acid metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | hypoglycin biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | felinine and 3-methyl-3-sulfanylbutan-1-ol biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | pau01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | glutathione degradation (DUG pathway - yeast) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | 4-hydroxy-2-nonenal detoxification | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF56235 | N-terminal nucleophile aminohydrolases (Ntn hydrolases) | IPR029055 | Nucleophile aminohydrolases, N-terminal | 34 | 564 | 0.0 |
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | - | - | 443 | 458 | 8.0E-58 |
| NCBIfam | TIGR00066 | JCVI: gamma-glutamyltransferase | IPR000101 | Gamma-glutamyltranspeptidase | 41 | 559 | 2.7E-108 |
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | - | - | 251 | 267 | 8.0E-58 |
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | - | - | 378 | 396 | 8.0E-58 |
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | - | - | 282 | 301 | 8.0E-58 |
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | - | - | 466 | 483 | 8.0E-58 |
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | - | - | 135 | 153 | 8.0E-58 |
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | - | - | 61 | 86 | 8.0E-58 |
| PANTHER | PTHR43199 | GLUTATHIONE HYDROLASE | - | - | 25 | 564 | 0.0 |
| Gene3D | G3DSA:1.10.246.130 | - | IPR043138 | Gamma-glutamyltranspeptidase, large subunit, C-terminal domain | 267 | 372 | 1.1E-24 |
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | - | - | 402 | 420 | 8.0E-58 |
| Pfam | PF01019 | Gamma-glutamyltranspeptidase | - | - | 54 | 561 | 0.0 |
| Gene3D | G3DSA:3.60.20.40 | - | IPR043137 | Gamma-glutamyltranspeptidase, small subunit | 378 | 566 | 2.2E-53 |
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | - | - | 153 | 172 | 8.0E-58 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.