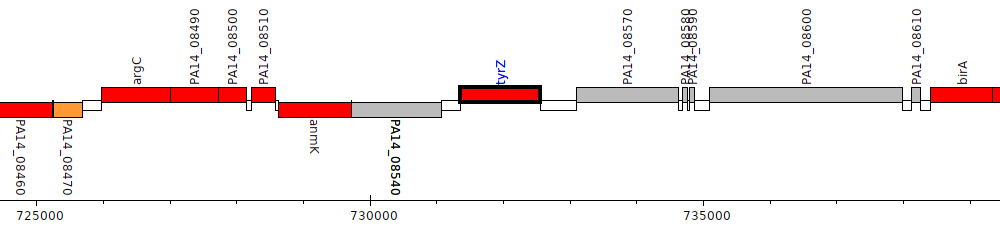
Pseudomonas aeruginosa UCBPP-PA14, PA14_08560 (tyrZ)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004812 | aminoacyl-tRNA ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00579
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00579
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000166 | nucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00579
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004831 | tyrosine-tRNA ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00234
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003723 | RNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.10.290.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006437 | tyrosyl-tRNA aminoacylation |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00234
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006418 | tRNA aminoacylation for protein translation |
Inferred from Sequence Model
Term mapped from: InterPro:PF00579
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pau00970 | Aminoacyl-tRNA biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd00805 | TyrRS_core | IPR002307 | Tyrosine-tRNA ligase | 34 | 298 | 8.29218E-115 |
| PANTHER | PTHR11766 | TYROSYL-TRNA SYNTHETASE | IPR024088 | Tyrosine-tRNA ligase, bacterial-type | 11 | 392 | 1.8E-87 |
| PRINTS | PR01040 | Tyrosyl-tRNA synthetase signature | IPR002307 | Tyrosine-tRNA ligase | 178 | 200 | 2.2E-26 |
| PRINTS | PR01040 | Tyrosyl-tRNA synthetase signature | IPR002307 | Tyrosine-tRNA ligase | 46 | 68 | 2.2E-26 |
| Hamap | MF_02007 | Tyrosine--tRNA ligase [tyrS]. | IPR024108 | Tyrosine-tRNA ligase, bacterial-type, type 2 | 2 | 396 | 48.201878 |
| FunFam | G3DSA:1.10.240.10:FF:000006 | Tyrosine--tRNA ligase | - | - | 219 | 325 | 2.9E-45 |
| PRINTS | PR01040 | Tyrosyl-tRNA synthetase signature | IPR002307 | Tyrosine-tRNA ligase | 211 | 223 | 2.2E-26 |
| SUPERFAMILY | SSF55174 | Alpha-L RNA-binding motif | - | - | 320 | 396 | 9.68E-14 |
| NCBIfam | TIGR00234 | JCVI: tyrosine--tRNA ligase | IPR002307 | Tyrosine-tRNA ligase | 2 | 396 | 5.3E-119 |
| FunFam | G3DSA:3.40.50.620:FF:000061 | Tyrosine--tRNA ligase | - | - | 2 | 218 | 1.6E-124 |
| PRINTS | PR01040 | Tyrosyl-tRNA synthetase signature | IPR002307 | Tyrosine-tRNA ligase | 161 | 176 | 2.2E-26 |
| Gene3D | G3DSA:1.10.240.10 | - | - | - | 219 | 322 | 9.8E-37 |
| SUPERFAMILY | SSF52374 | Nucleotidylyl transferase | - | - | 4 | 310 | 2.43E-99 |
| Gene3D | G3DSA:3.40.50.620 | HUPs | IPR014729 | Rossmann-like alpha/beta/alpha sandwich fold | 2 | 218 | 9.0E-83 |
| Pfam | PF00579 | tRNA synthetases class I (W and Y) | IPR002305 | Aminoacyl-tRNA synthetase, class Ic | 31 | 315 | 9.6E-81 |
| Gene3D | G3DSA:3.10.290.10 | - | IPR036986 | RNA-binding S4 domain superfamily | 323 | 387 | 6.7E-11 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.