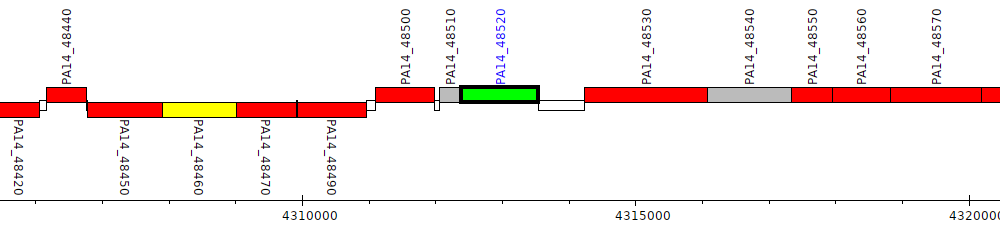
Pseudomonas aeruginosa UCBPP-PA14, PA14_48520
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0019867 | outer membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PF06725
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009254 | peptidoglycan turnover |
Inferred from Sequence Model
Term mapped from: InterPro:PF06725
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds |
Inferred from Sequence Model
Term mapped from: InterPro:SM00925
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM00925 | MltA_2 | IPR005300 | Lytic transglycosylase MltA, domain B | 126 | 281 | 4.4E-85 |
| Pfam | PF03562 | MltA specific insert domain | IPR005300 | Lytic transglycosylase MltA, domain B | 55 | 280 | 1.7E-75 |
| PANTHER | PTHR30124 | MEMBRANE-BOUND LYTIC MUREIN TRANSGLYCOSYLASE A | IPR026044 | Membrane-bound lytic murein transglycosylase A | 8 | 381 | 8.3E-127 |
| Gene3D | G3DSA:2.40.240.50 | - | - | - | 125 | 286 | 1.4E-108 |
| PIRSF | PIRSF019422 | MltA | IPR026044 | Membrane-bound lytic murein transglycosylase A | 2 | 383 | 1.0E-118 |
| SUPERFAMILY | SSF50685 | Barwin-like endoglucanases | IPR036908 | RlpA-like domain superfamily | 41 | 380 | 3.49E-125 |
| CDD | cd14668 | mlta_B | - | - | 127 | 283 | 1.33145E-74 |
| CDD | cd14485 | mltA_like_LT_A | - | - | 251 | 375 | 4.57442E-43 |
| Pfam | PF06725 | 3D domain | IPR010611 | 3D domain | 303 | 375 | 1.3E-23 |
| Gene3D | G3DSA:2.40.40.10 | - | IPR036908 | RlpA-like domain superfamily | 47 | 377 | 1.4E-108 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.