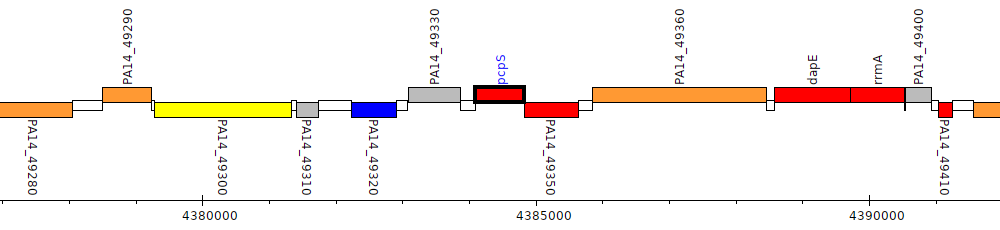
Pseudomonas aeruginosa UCBPP-PA14, PA14_49340 (pcpS)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016740 | transferase activity |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA1165
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
12381736 | |
| Molecular Function | GO:0000287 | magnesium ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF56214
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009239 | enterobactin biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR38096
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0009366 | enterobactin synthetase complex |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR38096
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008897 | holo-[acyl-carrier-protein] synthase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF56214
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016780 | phosphotransferase activity, for other substituted phosphate groups |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR38096
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Fatty acid biosynthesis (path 1); Pyochelin synthesis; Pyoverdine synthesis |
ECO:0000037
not_recorded |
|||
| KEGG | pau01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01053 | Biosynthesis of siderophore group nonribosomal peptides | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF01648 | 4'-phosphopantetheinyl transferase superfamily | IPR008278 | 4'-phosphopantetheinyl transferase domain | 125 | 220 | 2.4E-9 |
| SUPERFAMILY | SSF56214 | 4'-phosphopantetheinyl transferase | IPR037143 | 4'-phosphopantetheinyl transferase domain superfamily | 123 | 221 | 5.41E-13 |
| PANTHER | PTHR38096 | ENTEROBACTIN SYNTHASE COMPONENT D | IPR003542 | Enterobactin synthetase-like, component D | 1 | 240 | 9.8E-79 |
| PRINTS | PR01399 | Enterobactin synthetase component D signature | IPR003542 | Enterobactin synthetase-like, component D | 91 | 109 | 6.8E-18 |
| PRINTS | PR01399 | Enterobactin synthetase component D signature | IPR003542 | Enterobactin synthetase-like, component D | 61 | 81 | 6.8E-18 |
| PRINTS | PR01399 | Enterobactin synthetase component D signature | IPR003542 | Enterobactin synthetase-like, component D | 169 | 185 | 6.8E-18 |
| Pfam | PF17837 | 4'-phosphopantetheinyl transferase N-terminal domain | IPR041354 | 4'-phosphopantetheinyl transferase, N-terminal domain | 56 | 118 | 4.6E-21 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.