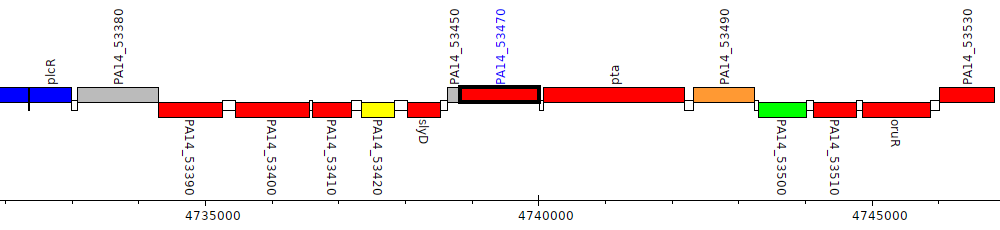
Pseudomonas aeruginosa UCBPP-PA14, PA14_53470
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016301 | kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000722
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016774 | phosphotransferase activity, carboxyl group as acceptor |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000722
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0016310 | phosphorylation |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR21060
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006082 | organic acid metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000722
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pau01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau00620 | Pyruvate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau00430 | Taurine and hypotaurine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01200 | Carbon metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau00640 | Propanoate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau00680 | Methane metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PIRSF | PIRSF000722 | Acetate_prop_kin | IPR004372 | Acetate/propionate kinase | 2 | 394 | 0.0 |
| PANTHER | PTHR21060 | ACETATE KINASE | IPR000890 | Aliphatic acid kinase, short-chain | 1 | 393 | 0.0 |
| PRINTS | PR00471 | Acetate kinase family signature | IPR000890 | Aliphatic acid kinase, short-chain | 319 | 335 | 6.4E-43 |
| SUPERFAMILY | SSF53067 | Actin-like ATPase domain | IPR043129 | ATPase, nucleotide binding domain | 153 | 393 | 1.11E-82 |
| Hamap | MF_00020 | Acetate kinase [ackA]. | IPR004372 | Acetate/propionate kinase | 1 | 394 | 169.520294 |
| Gene3D | G3DSA:3.30.420.40 | - | - | - | 2 | 192 | 3.9E-69 |
| PRINTS | PR00471 | Acetate kinase family signature | IPR000890 | Aliphatic acid kinase, short-chain | 6 | 17 | 6.4E-43 |
| Pfam | PF00871 | Acetokinase family | IPR000890 | Aliphatic acid kinase, short-chain | 6 | 386 | 0.0 |
| PRINTS | PR00471 | Acetate kinase family signature | IPR000890 | Aliphatic acid kinase, short-chain | 374 | 386 | 6.4E-43 |
| NCBIfam | TIGR00016 | JCVI: acetate/propionate family kinase | IPR004372 | Acetate/propionate kinase | 3 | 392 | 0.0 |
| SUPERFAMILY | SSF53067 | Actin-like ATPase domain | IPR043129 | ATPase, nucleotide binding domain | 5 | 192 | 5.83E-65 |
| PRINTS | PR00471 | Acetate kinase family signature | IPR000890 | Aliphatic acid kinase, short-chain | 200 | 221 | 6.4E-43 |
| PRINTS | PR00471 | Acetate kinase family signature | IPR000890 | Aliphatic acid kinase, short-chain | 297 | 310 | 6.4E-43 |
| PRINTS | PR00471 | Acetate kinase family signature | IPR000890 | Aliphatic acid kinase, short-chain | 170 | 183 | 6.4E-43 |
| Gene3D | G3DSA:3.30.420.40 | - | - | - | 196 | 394 | 1.1E-70 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.