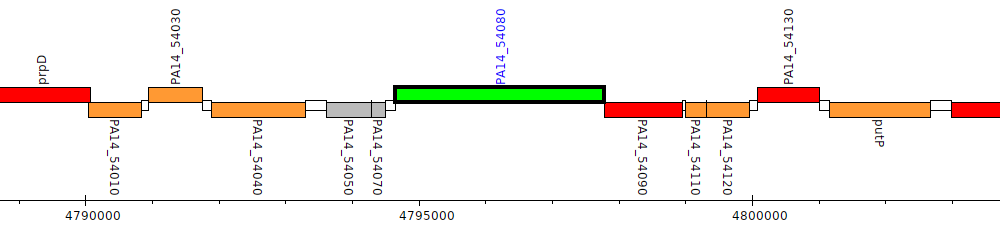
Pseudomonas aeruginosa UCBPP-PA14, PA14_54080
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | peptidoglycan biosynthesis III (mycobacteria) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | peptidoglycan biosynthesis IV (Enterococcus faecium) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | peptidoglycan biosynthesis V (β-lactam resistance) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | peptidoglycan biosynthesis II (staphylococci) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.710.10 | - | IPR012338 | Beta-lactamase/transpeptidase-like | 816 | 1023 | 1.1E-13 |
| Gene3D | G3DSA:1.10.3810.10 | - | IPR036950 | Penicillin binding protein transglycosylase domain | 163 | 397 | 3.2E-22 |
| Pfam | PF00912 | Transglycosylase | IPR001264 | Glycosyl transferase, family 51 | 162 | 359 | 8.6E-26 |
| PANTHER | PTHR32282 | BINDING PROTEIN TRANSPEPTIDASE, PUTATIVE-RELATED | - | - | 11 | 1021 | 2.3E-121 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 11 | 32 | - |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 1 | 33 | - |
| FunFam | G3DSA:3.40.710.10:FF:000047 | Glycosyl transferase family 51 | - | - | 815 | 1025 | 1.7E-124 |
| SUPERFAMILY | SSF56601 | beta-lactamase/transpeptidase-like | IPR012338 | Beta-lactamase/transpeptidase-like | 749 | 1023 | 1.52E-26 |
| SUPERFAMILY | SSF53955 | Lysozyme-like | IPR023346 | Lysozyme-like domain superfamily | 171 | 389 | 3.09E-9 |
| FunFam | G3DSA:1.10.3810.10:FF:000012 | Glycosyl transferase family 51 | - | - | 163 | 397 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.