Pseudomonas aeruginosa UCBPP-PA14, PA14_51340 (mvfR)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
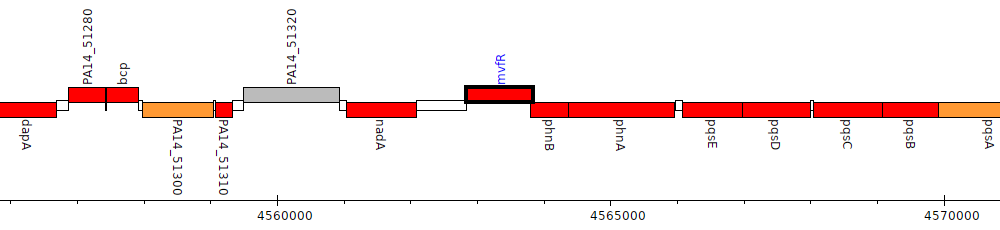
Gene Feature Overview
| Strain |
Pseudomonas aeruginosa UCBPP-PA14 (Lee et al., 2006)
GCF_000014625.1|latest |
| Locus Tag |
PA14_51340
|
| Name |
mvfR
|
| Replicon | chromosome |
| Genomic location | 4562837 - 4563835 (+ strand) |
| Transposon Mutants | 1 transposon mutants in UCBPP-PA14 |
| Transposon Mutants in orthologs | 3 transposon mutants in orthologs |
Cross-References
| RefSeq | YP_792268.1 |
| GI | 116048931 |
| Entrez | 4379987 |
| INSDC | ABJ10167.1 |
| NCBI Locus Tag | PA14_51340 |
| UniParc | UPI0000E44B70 |
| UniProtKB Acc | Q02IG8 |
| UniProtKB ID | Q02IG8_PSEAB |
| UniRef100 | UniRef100_Q02IG8 |
| UniRef50 | UniRef50_Q02IG8 |
| UniRef90 | UniRef90_Q02IG8 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
transcriptional regulator MvfR
|
| Synonyms | |
| Evidence for Translation | |
| Charge (pH 7) | 1.77 |
| Kyte-Doolittle Hydrophobicity Value | -0.131 |
| Molecular Weight (kDa) | 37.2 |
| Isoelectric Point (pI) | 7.56 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
PDB 3D Structures
| Accession | Header | Accession Date | Compound | Source | Resolution | Method | Percent Identity |
| 7P4U | GENE REGULATION | 07/13/21 | Crystal structure of PqsR (MvfR) ligand-binding domain in complex with 3-PYRIDIN-4-YL-2,4-DIHYDRO-INDENO[1,2-.C.]PYRAZOLE | Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) | 2.74 | X-RAY DIFFRACTION | 99.6 |
| 7NBW | DNA BINDING PROTEIN | 01/28/21 | Crystal structure of PqsR (MvfR) ligand-binding domain in complex with a pyridin agonist | Pseudomonas aeruginosa PAO1 | 2.28 | X-RAY DIFFRACTION | 99.6 |
| 6TPR | DNA BINDING PROTEIN | 12/14/19 | PqsR (MvfR) bound to inhibitory compound 40 | Pseudomonas aeruginosa UCBPP-PA14 | 3.2 | X-RAY DIFFRACTION | 100.0 |
| 7O2T | DNA BINDING PROTEIN | 03/31/21 | PqsR (MvfR) in complex with antagonist 61 | Pseudomonas aeruginosa | 2.65 | X-RAY DIFFRACTION | 100.0 |
| 7QAV | GENE REGULATION | 11/17/21 | Crystal structure of PqsR (MvfR) ligand-binding domain in complex with compound N-((2-(4-cyclopropylphenyl)thiazol-5-yl)methyl)-2-(trifluoromethyl)pyridin-4-amine | Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) | 2.65 | X-RAY DIFFRACTION | 99.4 |
| 4JVI | TRANSCRIPTION REGULATOR/INHIBITOR | 03/25/13 | Crystal structure of PqsR co-inducer binding domain of Pseudomonas aeruginosa with inhibitor 3NH2-7Cl-C9QZN | Pseudomonas aeruginosa | 2.9 | X-RAY DIFFRACTION | 100.0 |
| 6B8A | TRANSCRIPTION | 10/05/17 | Crystal structure of MvfR ligand binding domain in complex with M64 | Pseudomonas aeruginosa | 2.65 | X-RAY DIFFRACTION | 100.0 |
| 4JVD | TRANSCRIPTION REGULATOR | 03/25/13 | Crystal structure of PqsR coinducer binding domain of Pseudomonas aeruginosa with ligand NHQ | Pseudomonas aeruginosa | 2.95 | X-RAY DIFFRACTION | 100.0 |
| 6YIZ | DNA BINDING PROTEIN | 04/01/20 | Crystal structure of PqsR (MvfR) ligand-binding domain in complex with triazolo-pyridine inverse agonist A | Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) | 2.163 | X-RAY DIFFRACTION | 99.6 |
| 7QA3 | GENE REGULATION | 11/15/21 | Crystal structure of PqsR (MvfR) ligand-binding domain in complex with compound N-((2-(4-phenoxyphenyl)thiazol-5-yl)methyl)-2-(trifluoromethyl)pyridin-4-amine | Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) | 2.67 | X-RAY DIFFRACTION | 99.7 |
| 7QA0 | GENE REGULATION | 11/15/21 | Crystal structure of PqsR (MvfR) ligand-binding domain in complex with compound 1456 | Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) | 2.67 | X-RAY DIFFRACTION | 99.7 |
| 7O2U | DNA BINDING PROTEIN | 03/31/21 | PqsR (MvfR) in complex with antagonist 40 | Pseudomonas aeruginosa | 3 | X-RAY DIFFRACTION | 100.0 |
| 6Q7W | DNA BINDING PROTEIN | 12/13/18 | Crystal structure of PqsR (MvfR) ligand-binding domain in complex with compound 20 | Pseudomonas aeruginosa PAO1 | 2.82 | X-RAY DIFFRACTION | 99.6 |
| 6Q7U | DNA BINDING PROTEIN | 12/13/18 | Crystal structure of PqsR (MvfR) ligand-binding domain in complex with HHQ | Pseudomonas aeruginosa PAO1 | 3.14 | X-RAY DIFFRACTION | 99.6 |
| 6Z5K | DNA BINDING PROTEIN | 05/26/20 | PqsR (MvfR) in complex with antagonist 18 | Pseudomonas aeruginosa (strain UCBPP-PA14) | 3.201 | X-RAY DIFFRACTION | 100.0 |
| 6Z07 | DNA BINDING PROTEIN | 05/07/20 | PqsR (MvfR) in complex with antagonist 12 | Pseudomonas aeruginosa (strain UCBPP-PA14) | 2.95 | X-RAY DIFFRACTION | 100.0 |
| 6Q7V | DNA BINDING PROTEIN | 12/13/18 | Crystal structure of PqsR (MvfR) ligand-binding domain in complex with compound 11 | Pseudomonas aeruginosa PAO1 | 2.56 | X-RAY DIFFRACTION | 99.6 |
| 6Z17 | DNA BINDING PROTEIN | 05/12/20 | PqsR (MvfR) in complex with antagonist 6 | Pseudomonas aeruginosa (strain UCBPP-PA14) | 3.15 | X-RAY DIFFRACTION | 100.0 |
| 6YZ3 | DNA BINDING PROTEIN | 05/06/20 | PqsR (MvfR) in complex with antagonist 19 | Pseudomonas aeruginosa | 3 | X-RAY DIFFRACTION | 100.0 |
| 4JVC | TRANSCRIPTION REGULATOR | 03/25/13 | Crystal structure of PqsR co-inducer binding domain | Pseudomonas aeruginosa | 2.5 | X-RAY DIFFRACTION | 100.0 |
Virulence Evidence
| Source 1 | |
| Database | Victors : 4399 |
| Category | |
| Evidence |
ECO:0000015
mutant phenotype evidence
|
| Host Organism | Mus musculus |
| Notes | |
| Reference | 9371831 |
| Source 2 | |
| Database | PseudoCAP |
| Category | Phenazine biosynthesis |
| Evidence |
ECO:0000315
mutant phenotype evidence used in manual assertion
|
| Host Organism | Mus musculus |
| Notes | |
| Infection model | chronic pulmonary Infection model |
| Infection model | acute pulmonary Infection model |
| Reference | 15213173 |
| Source 3 | |
| Database | PseudoCAP |
| Category | |
| Evidence |
ECO:0000315
mutant phenotype evidence used in manual assertion
|
| Host Organism | Mus musculus |
| Notes | |
| Infection model | Mouse thermal injury model. |
| Reference | 11724939 |
| Source 4 | |
| Database | PseudoCAP |
| Category | |
| Evidence |
ECO:0000315
mutant phenotype evidence used in manual assertion
|
| Host Organism | Arabidopsis thaliana |
| Notes | |
| Infection model | Arabidopsis leaf infiltration assay |
| Reference | 11724939 |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 139 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG000966 (291 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PA14_51340
Search term: mvfR
Search term: transcriptional regulator MvfR
|
Human Homologs
References
|
A quorum sensing-associated virulence gene of Pseudomonas aeruginosa encodes a LysR-like transcription regulator with a unique self-regulatory mechanism.
Cao H, Krishnan G, Goumnerov B, Tsongalis J, Tompkins R, Rahme LG
Proc Natl Acad Sci U S A 2001 Dec 4;98(25):14613-8
PubMed ID: 11724939
|