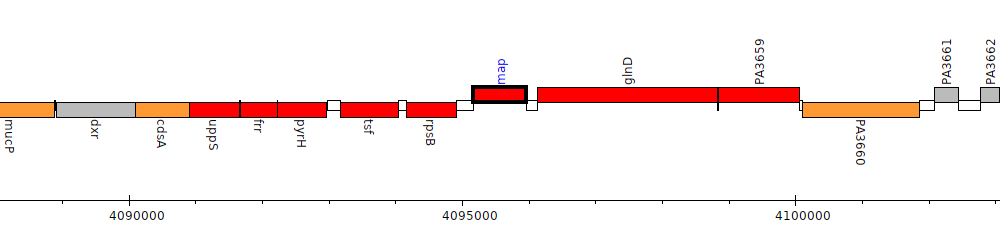
Pseudomonas aeruginosa PAO1, PA3657 (map)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Feature Overview
| Strain |
Pseudomonas aeruginosa PAO1 (Stover et al., 2000)
GCF_000006765.1|latest |
| Locus Tag |
PA3657
|
| Name |
map
|
| Replicon | chromosome |
| Genomic location | 4095172 - 4095957 (+ strand) |
| Transposon Mutants | 2 transposon mutants in PAO1 |
| Transposon Mutants in orthologs | 1 transposon mutants in orthologs |
Cross-References
| RefSeq | NP_252347.1 |
| GI | 15598853 |
| Affymetrix | PA3657_map_at |
| Entrez | 880564 |
| GenBank | AAG07045.1 |
| INSDC | AAG07045.1 |
| NCBI Locus Tag | PA3657 |
| protein_id(GenBank) | gb|AAG07045.1|AE004785_9|gnl|PseudoCAP|PA3657 |
| TIGR | NTL03PA03657 |
| UniParc | UPI00000C5AA5 |
| UniProtKB Acc | Q9HXY1 |
| UniProtKB ID | Q9HXY1_PSEAE |
| UniRef100 | UniRef100_W0YTW9 |
| UniRef50 | UniRef50_A6V1D1 |
| UniRef90 | UniRef90_A6V1D1 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
methionine aminopeptidase
|
| Synonyms |
peptidase M MAP |
| Evidence for Translation |
Identified using nanoflow high-pressure liquid chromatography (HPLC) in conjunction with microelectrospray ionization on LTQ XL mass spectrometer (PMID:24291602).
Detected by two-dimensional gel electrophoresis and mass spectral analysis(PMID:21863142).
|
| Charge (pH 7) | -4.53 |
| Kyte-Doolittle Hydrophobicity Value | -0.308 |
| Molecular Weight (kDa) | 29.1 |
| Isoelectric Point (pI) | 6.29 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
PDB 3D Structures
| Accession | Header | Accession Date | Compound | Source | Resolution | Method | Percent Identity |
| 4JUQ | HYDROLASE | 03/25/13 | Pseudomonas aeruginosa MetAP T2N mutant, in Mn form | Pseudomonas aeruginosa | 2.2 | X-RAY DIFFRACTION | 99.6 |
| 4FO8 | HYDROLASE | 06/20/12 | Pseudomonas aeruginosa MetAP with Met, in Mn form | Pseudomonas aeruginosa | 1.9 | X-RAY DIFFRACTION | 100.0 |
| 4FO7 | HYDROLASE | 06/20/12 | Pseudomonas aeruginosa MetAP, in Mn form | Pseudomonas aeruginosa | 1.8 | X-RAY DIFFRACTION | 100.0 |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 665 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG002017 (538 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PA3657
Search term: map
Search term: methionine aminopeptidase
|
Human Homologs
|
Ensembl
110, assembly
GRCh38.p14
|
methionyl aminopeptidase 1 [Source:HGNC Symbol;Acc:HGNC:15789]
E-value:
2.9e-75
Percent Identity:
52.0
|
References
|
Purification and characterization of a methionine-specific aminopeptidase from Salmonella typhimurium.
Wingfield P, Graber P, Turcatti G, Movva NR, Pelletier M, Craig S, Rose K, Miller CG
Eur J Biochem 1989 Mar 1;180(1):23-32
PubMed ID: 2651123
|
|
pepM is an essential gene in Salmonella typhimurium.
Miller CG, Kukral AM, Miller JL, Movva NR
J Bacteriol 1989 Sep;171(9):5215-7
PubMed ID: 2670909
|
|
Structure of the cobalt-dependent methionine aminopeptidase from Escherichia coli: a new type of proteolytic enzyme.
Roderick SL, Matthews BW
Biochemistry 1993 Apr 20;32(15):3907-12
PubMed ID: 8471602
|
|
Comparisons of Two Proteomic Analyses of Non-Mucoid and Mucoid Pseudomonas aeruginosa Clinical Isolates from a Cystic Fibrosis Patient.
Rao J, Damron FH, Basler M, Digiandomenico A, Sherman NE, Fox JW, Mekalanos JJ, Goldberg JB
Front Microbiol 2011;2:162
PubMed ID: 21863142
|