Pseudomonas aeruginosa PAO1, PA5489 (dsbA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
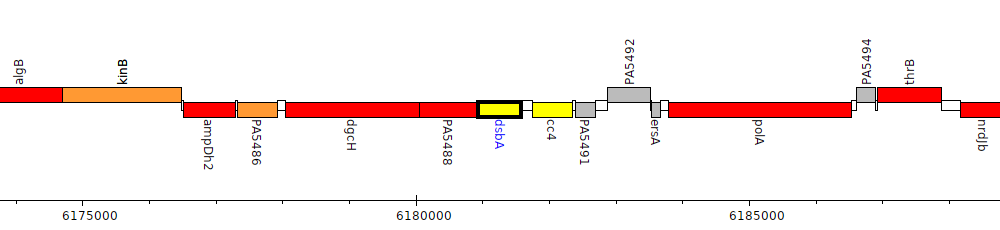
Gene Feature Overview
| Strain |
Pseudomonas aeruginosa PAO1 (Stover et al., 2000)
GCF_000006765.1|latest |
| Locus Tag |
PA5489
|
| Name |
dsbA
|
| Replicon | chromosome |
| Genomic location | 6180942 - 6181577 (- strand) |
| Transposon Mutants | 1 transposon mutants in PAO1 |
| Transposon Mutants in orthologs | 2 transposon mutants in orthologs |
Cross-References
| RefSeq | NP_254176.1 |
| GI | 15600682 |
| Affymetrix | PA5489_dsbA_at |
| DNASU | PaCD00008364 |
| Entrez | 877731 |
| GenBank | AAG08874.1 |
| INSDC | AAG08874.1 |
| NCBI Locus Tag | PA5489 |
| protein_id(GenBank) | gb|AAG08874.1|AE004961_9|gnl|PseudoCAP|PA5489 |
| TIGR | NTL03PA05490 |
| UniParc | UPI00001298C0 |
| UniProtKB Acc | P0C2B2 |
| UniProtKB ID | DSBA_PSEAE |
| UniRef100 | UniRef100_Q02DM0 |
| UniRef50 | UniRef50_O52376 |
| UniRef90 | UniRef90_Q02DM0 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
thiol:disulfide interchange protein DsbA
|
| Synonyms | |
| Evidence for Translation |
Detected by MALDI-TOF/TOF (PMID:19333994).
Identified using nanoflow high-pressure liquid chromatography (HPLC) in conjunction with microelectrospray ionization on LTQ XL mass spectrometer (PMID:24291602).
|
| Charge (pH 7) | -2.38 |
| Kyte-Doolittle Hydrophobicity Value | -0.064 |
| Molecular Weight (kDa) | 23.4 |
| Isoelectric Point (pI) | 6.40 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
PDB 3D Structures
| Accession | Header | Accession Date | Compound | Source | Resolution | Method | Percent Identity |
| 3H93 | TRANSCRIPTION REGULATOR | 04/29/09 | Crystal Structure of Pseudomonas aeruginosa DsbA | Pseudomonas aeruginosa PAO1 | 1.501 | X-RAY DIFFRACTION | 99.0 |
| 4ZL8 | OXIDOREDUCTASE | 05/01/15 | Crystal structure of Pseudomonas aeruginosa DsbA E82I: Crystal II | Pseudomonas aeruginosa | 1.395 | X-RAY DIFFRACTION | 98.4 |
| 5DCH | OXIDOREDUCTASE | 08/24/15 | Crystal structure of Pseudomonas aeruginosa DsbA E82I in complex with MIPS-0000851 (3-[(2-METHYLBENZYL)SULFANYL]-4H-1,2,4-TRIAZOL-4-AMINE) | Pseudomonas aeruginosa | 1.447 | X-RAY DIFFRACTION | 98.4 |
| 2MBT | OXIDOREDUCTASE | 08/03/13 | NMR study of PaDsbA | Pseudomonas aeruginosa | NOT | SOLUTION NMR | 100.0 |
| 4ZL7 | OXIDOREDUCTASE | 05/01/15 | Crystal structure of Pseudomonas aeruginosa DsbA E82I: Crystal I | Pseudomonas aeruginosa | 1.922 | X-RAY DIFFRACTION | 98.4 |
| 4ZL9 | OXIDOREDUCTASE | 05/01/15 | Crystal structure of Pseudomonas aeruginosa DsbA E82I: Crystal III | Pseudomonas aeruginosa | 1.7 | X-RAY DIFFRACTION | 98.4 |
| 5TLQ | OXIDOREDUCTASE | 10/11/16 | Model structure of the oxidized PaDsbA1 and 3-[(2-methylbenzyl)sulfanyl]-4H-1,2,4-triazol-4-amine complex | Pseudomonas aeruginosa | NOT | SOLUTION NMR | 100.0 |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 149 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG005029 (534 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PA5489
Search term: dsbA
Search term: thiol:disulfide interchange protein DsbA
|
Human Homologs
References
|
Genome-wide identification of Pseudomonas aeruginosa exported proteins using a consensus computational strategy combined with a laboratory-based PhoA fusion screen.
Lewenza S, Gardy JL, Brinkman FS, Hancock RE
Genome Res 2005 Feb;15(2):321-9
PubMed ID: 15687295
|
|
Cloning and expression of the gene for a protein disulfide oxidoreductase from Azotobacter vinelandii: complementation of an Escherichia coli dsbA mutant strain.
Ng TC, Kwik JF, Maier RJ
Gene 1997 Mar 25;188(1):109-13
PubMed ID: 9099867
|
|
Analysis of the periplasmic proteome of Pseudomonas aeruginosa, a metabolically versatile opportunistic pathogen.
Imperi F, Ciccosanti F, Perdomo AB, Tiburzi F, Mancone C, Alonzi T, Ascenzi P, Piacentini M, Visca P, Fimia GM
Proteomics 2009 Apr;9(7):1901-15
PubMed ID: 19333994
|