Pseudomonas aeruginosa PAO1, PA0996 (pqsA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
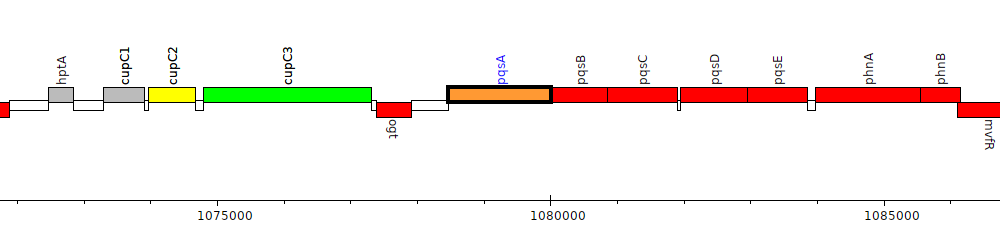
Gene Feature Overview
| Strain |
Pseudomonas aeruginosa PAO1 (Stover et al., 2000)
GCF_000006765.1|latest |
| Locus Tag |
PA0996
|
| Name |
pqsA
|
| Replicon | chromosome |
| Genomic location | 1078462 - 1080015 (+ strand) |
| Transposon Mutants | 2 transposon mutants in PAO1 |
| Transposon Mutants in orthologs | 1 transposon mutants in orthologs |
| Comment | Pseudomonas quinolone signal response |
Cross-References
| RefSeq | NP_249687.1 |
| GI | 15596193 |
| Affymetrix | PA0996_at |
| Entrez | 880760 |
| GenBank | AAG04385.1 |
| INSDC | AAG04385.1 |
| NCBI Locus Tag | PA0996 |
| protein_id(GenBank) | gb|AAG04385.1|AE004532_9|gnl|PseudoCAP|PA0996 |
| TIGR | NTL03PA00997 |
| UniParc | UPI00000C51F8 |
| UniProtKB Acc | Q9I4X3 |
| UniProtKB ID | PQSA_PSEAE |
| UniRef100 | UniRef100_Q9I4X3 |
| UniRef50 | UniRef50_Q9I4X3 |
| UniRef90 | UniRef90_Q9I4X3 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
PqsA
|
| Synonyms |
probable coenzyme A ligase |
| Evidence for Translation |
Identified using nanoflow high-pressure liquid chromatography (HPLC) in conjunction with microelectrospray ionization on LTQ XL mass spectrometer (PMID:24291602).
|
| Charge (pH 7) | -6.98 |
| Kyte-Doolittle Hydrophobicity Value | -0.068 |
| Molecular Weight (kDa) | 56.6 |
| Isoelectric Point (pI) | 6.15 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
PDB 3D Structures
| Accession | Header | Accession Date | Compound | Source | Resolution | Method | Percent Identity |
| 5OE3 | LIGASE | 07/07/17 | Crystal structure of the N-terminal domain of PqsA in complex with anthraniloyl-AMP (crystal form 1) | Pseudomonas aeruginosa PAO1 | 1.43 | X-RAY DIFFRACTION | 100.0 |
| 5OE5 | LIGASE | 07/07/17 | Crystal structure of the N-terminal domain of PqsA in complex with anthraniloyl-AMP (crystal form 3) | Pseudomonas aeruginosa PAO1 | 1.74 | X-RAY DIFFRACTION | 100.0 |
| 5OE6 | LIGASE | 07/07/17 | Crystal structure of the N-terminal domain of PqsA in complex with 6-fluoroanthraniloyl-AMP (crystal form 1) | Pseudomonas aeruginosa PAO1 | 1.67 | X-RAY DIFFRACTION | 100.0 |
| 5OE4 | LIGASE | 07/07/17 | Crystal structure of the N-terminal domain of PqsA in complex with anthraniloyl-AMP (crystal form 2) | Pseudomonas aeruginosa PAO1 | 1.904 | X-RAY DIFFRACTION | 100.0 |
Virulence Evidence
| Source 1 | |
| Database | PseudoCAP |
| Category | Not available / Not known |
| Evidence |
ECO:0000315
mutant phenotype evidence used in manual assertion
|
| Host Organism | Populus trichocarpa |
| Notes | |
| Infection model | Poplar pathogenicity wilting assay |
| Reference | 21261818 |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 600 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG000959 (296 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PA0996
Search term: pqsA
Search term: PqsA
|
Human Homologs
|
Ensembl
110, assembly
GRCh38.p14
|
acyl-CoA synthetase short chain family member 1 [Source:HGNC Symbol;Acc:HGNC:16091]
E-value:
1.6e-24
Percent Identity:
24.3
|
|
Ensembl
110, assembly
GRCh38.p14
|
acyl-CoA synthetase short chain family member 1 [Source:HGNC Symbol;Acc:HGNC:16091]
E-value:
1.6e-24
Percent Identity:
24.3
|
References
|
Genome-wide identification of Pseudomonas aeruginosa exported proteins using a consensus computational strategy combined with a laboratory-based PhoA fusion screen.
Lewenza S, Gardy JL, Brinkman FS, Hancock RE
Genome Res 2005 Feb;15(2):321-9
PubMed ID: 15687295
|
|
4-Hydroxybenzoate-coenzyme A ligase from Rhodopseudomonas palustris: purification, gene sequence, and role in anaerobic degradation.
Gibson J, Dispensa M, Fogg GC, Evans DT, Harwood CS
J Bacteriol 1994 Feb;176(3):634-41
PubMed ID: 8300518
|
|
PqsA is required for the biosynthesis of 2,4-dihydroxyquinoline (DHQ), a newly identified metabolite produced by Pseudomonas aeruginosa and Burkholderia thailandensis.
Lépine F, Dekimpe V, Lesic B, Milot S, Lesimple A, Mamer OA, Rahme LG, Déziel E
Biol Chem 2007 Aug;388(8):839-45
PubMed ID: 17655503
|