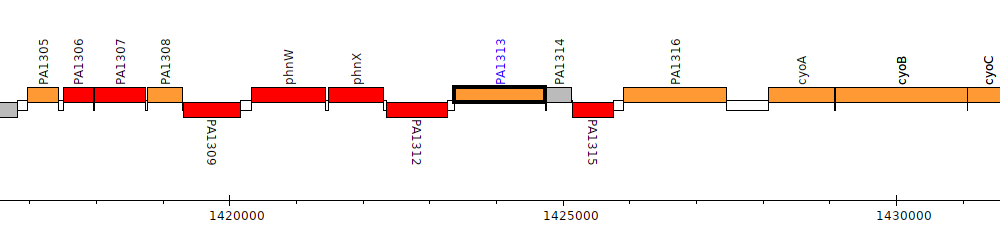
Pseudomonas aeruginosa PAO1, PA1313
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Feature Overview
| Strain |
Pseudomonas aeruginosa PAO1 (Stover et al., 2000)
GCF_000006765.1|latest |
| Locus Tag |
PA1313
|
| Name |
|
| Replicon | chromosome |
| Genomic location | 1423374 - 1424732 (+ strand) |
| Transposon Mutants | 4 transposon mutants in PAO1 |
| Transposon Mutants in orthologs | 1 transposon mutants in orthologs |
Cross-References
| RefSeq | NP_250004.1 |
| GI | 15596510 |
| Affymetrix | PA1313_at |
| Entrez | 881630 |
| GenBank | AAG04702.1 |
| INSDC | AAG04702.1 |
| NCBI Locus Tag | PA1313 |
| protein_id(GenBank) | gb|AAG04702.1|AE004560_10|gnl|PseudoCAP|PA1313 |
| TIGR | NTL03PA01314 |
| UniParc | UPI00000C5308 |
| UniProtKB Acc | Q9I431 |
| UniProtKB ID | Q9I431_PSEAE |
| UniRef100 | UniRef100_U9HQA2 |
| UniRef50 | UniRef50_D4E2K7 |
| UniRef90 | UniRef90_B7UVR3 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
probable major facilitator superfamily (MFS) transporter
|
| Synonyms |
probable MFS transporter |
| Evidence for Translation | |
| Charge (pH 7) | 20.67 |
| Kyte-Doolittle Hydrophobicity Value | 0.840 |
| Molecular Weight (kDa) | 47.2 |
| Isoelectric Point (pI) | 11.15 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 485 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG001256 (359 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PA1313
|
Human Homologs
References
|
Genome-wide identification of Pseudomonas aeruginosa exported proteins using a consensus computational strategy combined with a laboratory-based PhoA fusion screen.
Lewenza S, Gardy JL, Brinkman FS, Hancock RE
Genome Res 2005 Feb;15(2):321-9
PubMed ID: 15687295
|
|
Sequence and transcriptional analysis of the Streptomyces glaucescens tcmAR tetracenomycin C resistance and repressor gene loci.
Guilfoile PG, Hutchinson CR
J Bacteriol 1992 Jun;174(11):3651-8
PubMed ID: 1592819
|
|
Cloning, sequencing and deduced functions of a cluster of Streptomyces genes probably encoding biosynthesis of the polyketide antibiotic frenolicin.
Bibb MJ, Sherman DH, Omura S, Hopwood DA
Gene 1994 May 3;142(1):31-9
PubMed ID: 8181754
|
|
EmrR is a negative regulator of the Escherichia coli multidrug resistance pump EmrAB.
Lomovskaya O, Lewis K, Matin A
J Bacteriol 1995 May;177(9):2328-34
PubMed ID: 7730261
|