Pseudomonas aeruginosa PAO1, PA1314
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Feature Overview
| Strain |
Pseudomonas aeruginosa PAO1 (Stover et al., 2000)
GCF_000006765.1|latest |
| Locus Tag |
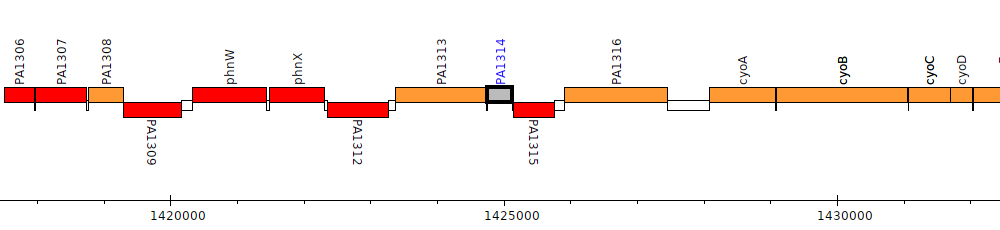
PA1314
|
| Name |
|
| Replicon | chromosome |
| Genomic location | 1424746 - 1425132 (+ strand) |
| Transposon Mutants | 2 transposon mutants in PAO1 |
| Transposon Mutants in orthologs | 1 transposon mutants in orthologs |
Cross-References
| RefSeq | NP_250005.1 |
| GI | 15596511 |
| Affymetrix | PA1314_at |
| DNASU | PaCD00006606 |
| Entrez | 881529 |
| GenBank | AAG04703.1 |
| INSDC | AAG04703.1 |
| NCBI Locus Tag | PA1314 |
| protein_id(GenBank) | gb|AAG04703.1|AE004560_11|gnl|PseudoCAP|PA1314 |
| TIGR | NTL03PA01315 |
| UniParc | UPI00000C5309 |
| UniProtKB Acc | Q9I430 |
| UniProtKB ID | Q9I430_PSEAE |
| UniRef100 | UniRef100_W0YP09 |
| UniRef50 | UniRef50_W0YP09 |
| UniRef90 | UniRef90_W0YP09 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
hypothetical protein
|
| Synonyms | |
| Evidence for Translation | |
| Charge (pH 7) | -2.63 |
| Kyte-Doolittle Hydrophobicity Value | -0.297 |
| Molecular Weight (kDa) | 14.2 |
| Isoelectric Point (pI) | 6.23 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
PDB 3D Structures
| Accession | Header | Accession Date | Compound | Source | Resolution | Method | Percent Identity |
| 1TP6 | structural genomics, unknown function | 06/15/04 | 1.5 A Crystal Structure of a NTF-2 Like Protein of Unknown Function PA1314 from Pseudomonas aeruginosa | Pseudomonas aeruginosa PAO1 | 1.5 | X-RAY DIFFRACTION | 100.0 |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 25 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG001257 (308 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PA1314
|
Human Homologs
References
No references are associated with this feature.