Pseudomonas putida BIRD-1, PPUBIRD1_3225
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
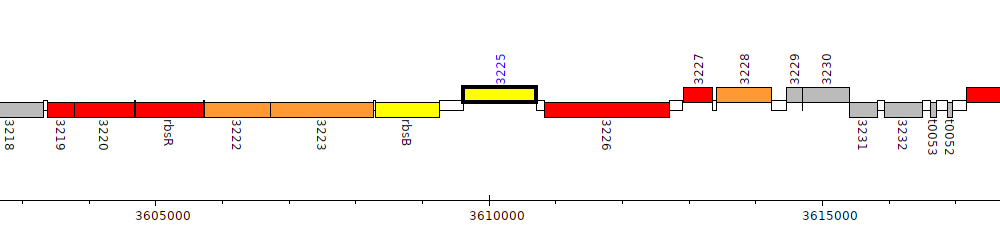
Gene Feature Overview
| Strain |
Pseudomonas putida BIRD-1
GCF_000183645.1|latest |
| Locus Tag |
PPUBIRD1_3225
|
| Name |
|
| Replicon | chromosome |
| Genomic location | 3609619 - 3610707 (+ strand) |
Cross-References
| RefSeq | YP_005931029.1 |
| GI | 386012752 |
| EC Number | 3.5.1.38 |
| Entrez | 12521394 |
| NCBI Locus Tag | PPUBIRD1_3225 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
Glutaminase-asparaginase
|
| Synonyms | |
| Evidence for Translation | |
| Charge (pH 7) | 1.21 |
| Kyte-Doolittle Hydrophobicity Value | -0.108 |
| Molecular Weight (kDa) | 38.6 |
| Isoelectric Point (pI) | 7.84 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
AlphaFold 2 Protein Structure Predictions
Protein structure predictions using a neural network model developed by DeepMind. If a UniProtKB accession is associated with this protein, a search link will be provided below.
PDB 3D Structures
| Accession | Header | Accession Date | Compound | Source | Resolution | Method | Percent Identity |
| 6WYW | HYDROLASE | 05/13/20 | Crystal structure of Pseudomonas 7A Glutaminase-Asparaginase in complex with L-Asp at pH 4.5 | Pseudomonas putida (strain ATCC 47054 / DSM 6125 / NCIMB 11950 / KT2440) | 2.13 | X-RAY DIFFRACTION | 99.7 |
| 6WYY | HYDROLASE | 05/13/20 | Crystal structure of Pseudomonas 7A Glutaminase-Asparaginase in complex with L-Glu at pH 6.5 | Pseudomonas putida (strain ATCC 47054 / DSM 6125 / NCIMB 11950 / KT2440) | 1.35 | X-RAY DIFFRACTION | 99.7 |
| 6WZ4 | HYDROLASE | 05/13/20 | Complex of mutant (K173M) of Pseudomonas 7A Glutaminase-Asparaginase with L-Asp at pH 6. Covalent acyl-enzyme intermediate | Pseudomonas putida (strain ATCC 47054 / DSM 6125 / NCIMB 11950 / KT2440) | 1.58 | X-RAY DIFFRACTION | 99.4 |
| 6WZ8 | HYDROLASE | 05/13/20 | Complex of Pseudomonas 7A Glutaminase-Asparaginase with Citrate Anion at pH 5.5. Covalent Acyl-Enzyme Intermediate | Pseudomonas putida (strain ATCC 47054 / DSM 6125 / NCIMB 11950 / KT2440) | 1.7 | X-RAY DIFFRACTION | 99.7 |
| 6WYX | HYDROLASE | 05/13/20 | Crystal structure of Pseudomonas 7A Glutaminase-Asparaginase in complex with L-Asp at pH 5.0 | Pseudomonas putida (strain ATCC 47054 / DSM 6125 / NCIMB 11950 / KT2440) | 1.48 | X-RAY DIFFRACTION | 99.7 |
| 6WZ6 | HYDROLASE | 05/13/20 | Complex of mutant (K173M) of Pseudomonas 7A Glutaminase-Asparaginase with L-Glu at pH 5. Covalent acyl-enzyme intermediate | Pseudomonas putida (strain ATCC 47054 / DSM 6125 / NCIMB 11950 / KT2440) | 1.15 | X-RAY DIFFRACTION | 99.4 |
| 6WYZ | HYDROLASE | 05/13/20 | Crystal structure of Pseudomonas 7A Glutaminase-Asparaginase (mutant K173M) in complex with D-Glu at pH 5.5 | Pseudomonas putida (strain ATCC 47054 / DSM 6125 / NCIMB 11950 / KT2440) | 1.7 | X-RAY DIFFRACTION | 99.4 |
Drugs Targeting this Protein
Identified by Diamond using e-value cutoff of 0.0001 and returning alignments that span 100% of the query sequence and that have more than 95% identity.
| Drug Name | Source Accession | Source DB | Version | Target Accession | Target Description | Percent Identity | Alignment Length | E-Value |
| Threonine-Aspartic Ester | DB01817 | DrugBank | 5.1.4 | P00805 | L-asparaginase 2 | 45.4 | 359 | 2.6e-74 |
| 2-Amino-6-Oxo-Hexanoic Acid | DB02571 | DrugBank | 5.1.4 | Q88K39 | Glutaminase-asparaginase | 99.7 | 362 | 1.4e-195 |
| 4-Carboxy-4-Aminobutanal | DB04388 | DrugBank | 5.1.4 | Q88K39 | Glutaminase-asparaginase | 99.7 | 362 | 1.4e-195 |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 351 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG001273 (453 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PPUBIRD1_3225
Search term: Glutaminase-asparaginase
|
Human Homologs
References
No references are associated with this feature.