Pseudomonas aeruginosa M18, PAM18_3913
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Feature Overview
| Strain |
Pseudomonas aeruginosa M18
GCF_000226155.1|latest |
| Locus Tag |
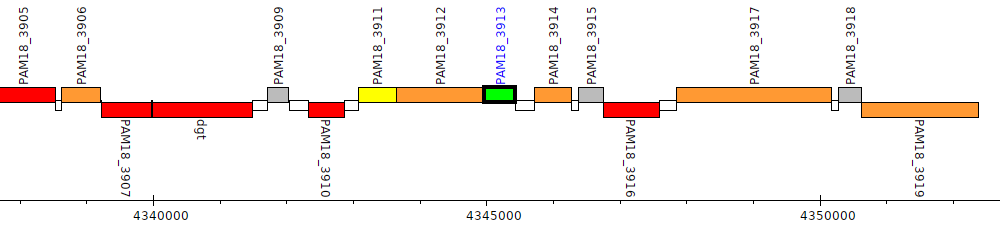
PAM18_3913
|
| Name |
|
| Replicon | chromosome |
| Genomic location | 4344962 - 4345438 (+ strand) |
Cross-References
| RefSeq | YP_005976496.1 |
| GI | 386059974 |
| Entrez | 12567967 |
| NCBI Locus Tag | PAM18_3913 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
putative outer membrane protein
|
| Synonyms | |
| Evidence for Translation | |
| Charge (pH 7) | -0.79 |
| Kyte-Doolittle Hydrophobicity Value | -0.297 |
| Molecular Weight (kDa) | 17.2 |
| Isoelectric Point (pI) | 5.98 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
AlphaFold 2 Protein Structure Predictions
Protein structure predictions using a neural network model developed by DeepMind. If a UniProtKB accession is associated with this protein, a search link will be provided below.
PDB 3D Structures
| Accession | Header | Accession Date | Compound | Source | Resolution | Method | Percent Identity |
| 5Y61 | SIGNALING PROTEIN | 08/10/17 | YfiB-YfiR complexed with GMP | Pseudomonas aeruginosa | 2.99 | X-RAY DIFFRACTION | 99.3 |
| 5EB0 | MEMBRANE PROTEIN | 10/17/15 | crystal form II of YfiB belonging to P41 | Pseudomonas aeruginosa PAO1 | 2.8 | X-RAY DIFFRACTION | 100.0 |
| 5EAZ | MEMBRANE PROTEIN | 10/17/15 | crystal form I of YfiB belonging to space groups P21 | Pseudomonas aeruginosa PAO1 | 2.151 | X-RAY DIFFRACTION | 100.0 |
| 4ZHY | TRANSCRIPTION/SIGNALING PROTEIN | 04/27/15 | Crystal structure of a bacterial signalling complex | Pseudomonas aeruginosa PAO1 | 1.969 | X-RAY DIFFRACTION | 97.5 |
| 6IKI | SIGNALING PROTEIN | 10/16/18 | Crystal structure of YfiB(W55L) | Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) | 2.204 | X-RAY DIFFRACTION | 98.7 |
| 6IKJ | SIGNALING PROTEIN | 10/16/18 | Crystal structure of YfiB(F48S) | Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) | 1.76 | X-RAY DIFFRACTION | 98.7 |
| 6IKK | SIGNALING PROTEIN | 10/16/18 | Crystal structure of YfiB(L43P) in complex with YfiR | Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) | 2.19 | X-RAY DIFFRACTION | 98.7 |
| 4ZHW | SIGNALING PROTEIN | 04/27/15 | Crystal structure of a bacterial signalling protein (N-terminal truncation) | Pseudomonas aeruginosa PAO1 | 1.391 | X-RAY DIFFRACTION | 99.4 |
| 5EB1 | MEMBRANE PROTEIN/TRANSCRIPTION | 10/17/15 | the YfiB-YfiR complex | Pseudomonas aeruginosa PAO1 | 1.8 | X-RAY DIFFRACTION | 99.3 |
| 4ZHV | SIGNALING PROTEIN | 04/27/15 | Crystal structure of a bacterial signalling protein | Pseudomonas aeruginosa PAO1 | 1.585 | X-RAY DIFFRACTION | 99.4 |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 408 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG001079 (445 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PAM18_3913
Search term: putative outer membrane protein
|
Human Homologs
References
No references are associated with this feature.