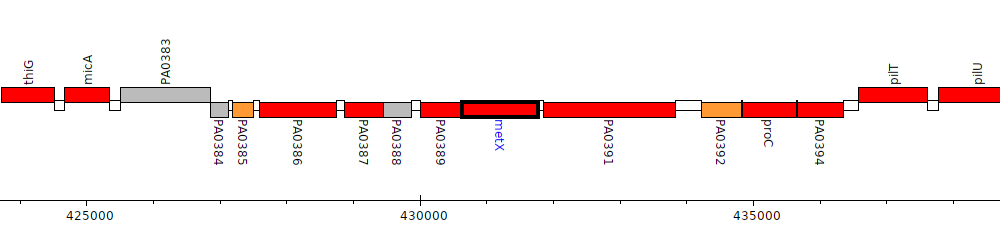
Pseudomonas aeruginosa PAO1, PA0390 (metX)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0008137 | NADH dehydrogenase (ubiquinone) activity | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006790 | sulfur compound metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006520 | cellular amino acid metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0016747 | transferase activity, transferring acyl groups other than amino-acyl groups |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR32268
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009058 | biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR32268
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCyc | PWY-5344 | L-homocysteine biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | PWY-821 | superpathway of sulfur amino acid biosynthesis (Saccharomyces cerevisiae) | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | HSERMETANA-PWY | L-methionine biosynthesis III | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00270 | Cysteine and methionine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Sulfur metabolism |
ECO:0000037
not_recorded |
|||
| PseudoCAP | Methionine metabolism |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PIRSF | PIRSF000443 | Homoser_Ac_trans | IPR008220 | Homoserine/serine acetyltransferase MetX-like | 2 | 379 | 0.0 |
| PANTHER | PTHR32268 | HOMOSERINE O-ACETYLTRANSFERASE | IPR008220 | Homoserine/serine acetyltransferase MetX-like | 14 | 374 | 9.4E-128 |
| SUPERFAMILY | SSF53474 | alpha/beta-Hydrolases | IPR029058 | Alpha/Beta hydrolase fold | 13 | 375 | 9.76E-101 |
| Pfam | PF00561 | alpha/beta hydrolase fold | IPR000073 | Alpha/beta hydrolase fold-1 | 51 | 360 | 5.5E-32 |
| Gene3D | G3DSA:1.10.1740.110 | - | - | - | 187 | 293 | 0.0 |
| FunFam | G3DSA:1.10.1740.110:FF:000001 | Homoserine O-acetyltransferase | - | - | 187 | 293 | 2.7E-44 |
| NCBIfam | TIGR01392 | JCVI: homoserine O-acetyltransferase | IPR008220 | Homoserine/serine acetyltransferase MetX-like | 21 | 370 | 0.0 |
| Gene3D | G3DSA:3.40.50.1820 | alpha/beta hydrolase | IPR029058 | Alpha/Beta hydrolase fold | 26 | 361 | 0.0 |
| Hamap | MF_00296 | Homoserine O-acetyltransferase [metXA]. | IPR008220 | Homoserine/serine acetyltransferase MetX-like | 13 | 375 | 28.601257 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.