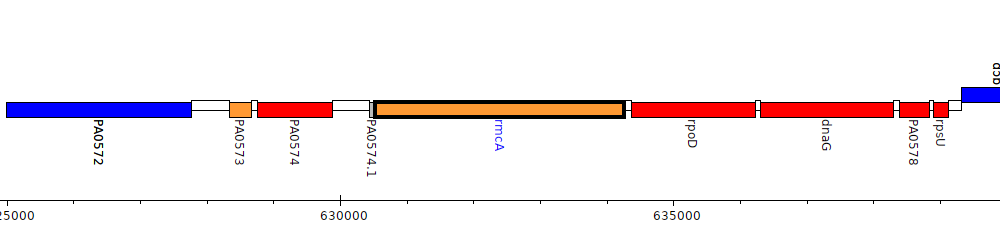
Pseudomonas aeruginosa PAO1, PA0575 (rmcA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0005515 | protein binding |
Term mapped from: PseudoCAP:PA4959
|
ECO:0007073 |
35732459 | Reviewed by curator |
| Molecular Function | GO:0005515 | protein binding |
Term mapped from: PseudoCAP:PA4601
|
ECO:0007073 |
35732459 | Reviewed by curator |
| Molecular Function | GO:0005515 | protein binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF08447
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF55785 | PYP-like sensor domain (PAS domain) | IPR035965 | PAS domain superfamily | 688 | 795 | 5.24E-29 |
| SUPERFAMILY | SSF141868 | EAL domain-like | IPR035919 | EAL domain superfamily | 984 | 1231 | 1.7E-89 |
| PANTHER | PTHR44757 | DIGUANYLATE CYCLASE DGCP | - | - | 665 | 1236 | 0.0 |
| NCBIfam | TIGR00229 | JCVI: PAS domain S-box protein | IPR000014 | PAS domain | 308 | 432 | 1.0E-13 |
| SMART | SM00091 | pas_2 | IPR000014 | PAS domain | 562 | 629 | 1.1E-6 |
| CDD | cd00130 | PAS | IPR000014 | PAS domain | 692 | 794 | 9.38124E-13 |
| Pfam | PF00563 | EAL domain | IPR001633 | EAL domain | 986 | 1220 | 7.0E-77 |
| NCBIfam | TIGR00229 | JCVI: PAS domain S-box protein | IPR000014 | PAS domain | 687 | 804 | 1.9E-19 |
| Gene3D | G3DSA:3.40.190.10 | - | - | - | 28 | 254 | 5.8E-69 |
| SUPERFAMILY | SSF53850 | Periplasmic binding protein-like II | - | - | 39 | 256 | 1.59E-49 |
| SMART | SM00062 | AABind_6 | IPR001638 | Solute-binding protein family 3/N-terminal domain of MltF | 38 | 258 | 2.2E-32 |
| SMART | SM00086 | pac_2 | IPR001610 | PAC motif | 755 | 797 | 5.5E-5 |
| Pfam | PF13426 | PAS domain | IPR000014 | PAS domain | 572 | 674 | 4.3E-8 |
| NCBIfam | TIGR00229 | JCVI: PAS domain S-box protein | IPR000014 | PAS domain | 436 | 556 | 2.0E-11 |
| Gene3D | G3DSA:3.20.20.450 | EAL domain | IPR035919 | EAL domain superfamily | 979 | 1245 | 7.8E-105 |
| CDD | cd00130 | PAS | IPR000014 | PAS domain | 331 | 423 | 1.06051E-6 |
| Pfam | PF00497 | Bacterial extracellular solute-binding proteins, family 3 | IPR001638 | Solute-binding protein family 3/N-terminal domain of MltF | 39 | 256 | 4.9E-36 |
| Pfam | PF13426 | PAS domain | IPR000014 | PAS domain | 448 | 548 | 1.3E-4 |
| SMART | SM00086 | pac_2 | IPR001610 | PAC motif | 386 | 426 | 1.0E-5 |
| SUPERFAMILY | SSF55073 | Nucleotide cyclase | IPR029787 | Nucleotide cyclase | 813 | 970 | 4.79E-54 |
| SUPERFAMILY | SSF55785 | PYP-like sensor domain (PAS domain) | IPR035965 | PAS domain superfamily | 297 | 424 | 8.38E-21 |
| CDD | cd00130 | PAS | IPR000014 | PAS domain | 572 | 672 | 8.27902E-8 |
| CDD | cd00130 | PAS | IPR000014 | PAS domain | 448 | 547 | 1.09193E-6 |
| FunFam | G3DSA:3.30.70.270:FF:000001 | Diguanylate cyclase domain protein | - | - | 803 | 970 | 7.4E-45 |
| SMART | SM00091 | pas_2 | IPR000014 | PAS domain | 683 | 749 | 6.9E-8 |
| SUPERFAMILY | SSF55785 | PYP-like sensor domain (PAS domain) | IPR035965 | PAS domain superfamily | 423 | 548 | 2.44E-19 |
| CDD | cd01948 | EAL | IPR001633 | EAL domain | 985 | 1224 | 2.67639E-114 |
| Gene3D | G3DSA:3.30.450.20 | PAS domain | - | - | 680 | 809 | 7.2E-37 |
| Pfam | PF08447 | PAS fold | IPR013655 | PAS fold-3 | 336 | 408 | 3.1E-10 |
| NCBIfam | TIGR00229 | JCVI: PAS domain S-box protein | IPR000014 | PAS domain | 560 | 682 | 7.8E-16 |
| CDD | cd01007 | PBP2_BvgS_HisK_like | - | - | 36 | 255 | 2.28936E-80 |
| Pfam | PF00990 | Diguanylate cyclase, GGDEF domain | IPR000160 | GGDEF domain | 808 | 965 | 1.3E-53 |
| SMART | SM00086 | pac_2 | IPR001610 | PAC motif | 635 | 675 | 83.0 |
| SMART | SM00091 | pas_2 | IPR000014 | PAS domain | 438 | 505 | 3.2E-6 |
| Gene3D | G3DSA:3.30.450.20 | PAS domain | - | - | 303 | 423 | 3.7E-23 |
| SMART | SM00267 | duf1_3 | IPR000160 | GGDEF domain | 797 | 969 | 2.1E-71 |
| SMART | SM00086 | pac_2 | IPR001610 | PAC motif | 510 | 550 | 3.5 |
| Gene3D | G3DSA:3.30.450.20 | PAS domain | - | - | 551 | 673 | 4.9E-20 |
| Gene3D | G3DSA:3.40.190.10 | - | - | - | 123 | 217 | 5.8E-69 |
| NCBIfam | TIGR00254 | JCVI: diguanylate cyclase (GGDEF) domain | IPR000160 | GGDEF domain | 804 | 968 | 5.5E-44 |
| SMART | SM00091 | pas_2 | IPR000014 | PAS domain | 312 | 380 | 0.0026 |
| Coils | Coil | Coil | - | - | 286 | 320 | - |
| CDD | cd01949 | GGDEF | IPR000160 | GGDEF domain | 809 | 966 | 2.46212E-70 |
| SUPERFAMILY | SSF55785 | PYP-like sensor domain (PAS domain) | IPR035965 | PAS domain superfamily | 547 | 673 | 1.64E-19 |
| Gene3D | G3DSA:3.30.70.270 | - | IPR043128 | Reverse transcriptase/Diguanylate cyclase domain | 810 | 978 | 1.0E-65 |
| FunFam | G3DSA:3.20.20.450:FF:000001 | Cyclic di-GMP phosphodiesterase yahA | - | - | 980 | 1235 | 5.7E-98 |
| Pfam | PF13426 | PAS domain | IPR000014 | PAS domain | 694 | 795 | 3.3E-15 |
| Gene3D | G3DSA:3.30.450.20 | PAS domain | - | - | 424 | 548 | 3.2E-21 |
| SMART | SM00052 | duf2_2 | IPR001633 | EAL domain | 979 | 1225 | 1.3E-123 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.