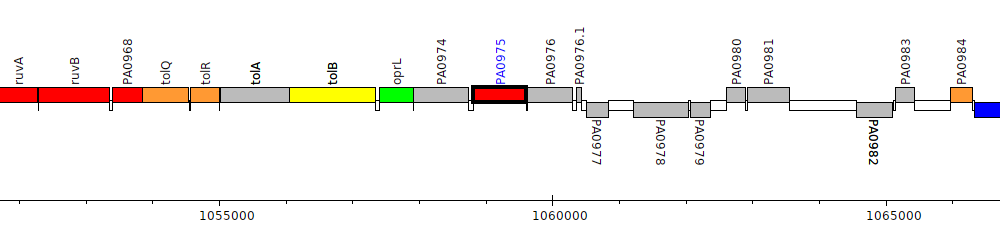
Pseudomonas aeruginosa PAO1, PA0975
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0051536 | iron-sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:SFLDS00029
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SFLDS00029
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00790 | Folate biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.20.20.70 | Aldolase class I | IPR013785 | Aldolase-type TIM barrel | 53 | 258 | 6.3E-43 |
| SFLD | SFLDS00029 | Radical SAM | IPR007197 | Radical SAM | 77 | 215 | 6.4E-20 |
| Pfam | PF04055 | Radical SAM superfamily | IPR007197 | Radical SAM | 77 | 197 | 4.3E-15 |
| CDD | cd01335 | Radical_SAM | - | - | 80 | 153 | 1.03721E-8 |
| SUPERFAMILY | SSF102114 | Radical SAM enzymes | - | - | 76 | 154 | 1.7E-14 |
| PANTHER | PTHR42836 | 7-CARBOXY-7-DEAZAGUANINE SYNTHASE | - | - | 50 | 263 | 1.0E-58 |
| NCBIfam | TIGR04349 | JCVI: 7-carboxy-7-deazaguanine synthase QueE | IPR027621 | Putative 7-cyano-7-deazaguanosine (preQ0) biosynthesis protein QueE | 54 | 264 | 2.1E-110 |
| Pfam | PF13394 | 4Fe-4S single cluster domain | - | - | 75 | 156 | 4.1E-10 |
| Hamap | MF_00917 | 7-carboxy-7-deazaguanine synthase [queE]. | IPR024924 | 7-carboxy-7-deazaguanine synthase-like | 54 | 261 | 35.976929 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.