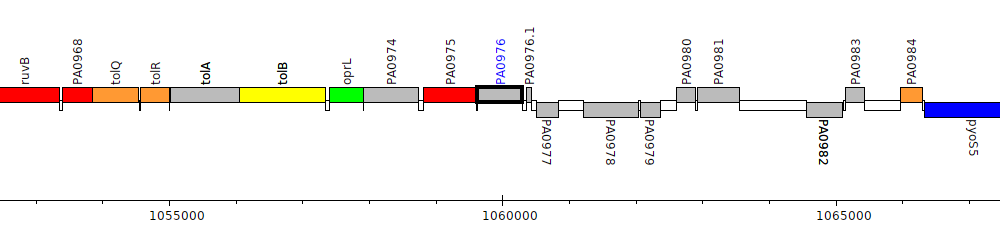
Pseudomonas aeruginosa PAO1, PA0976
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00790 | Folate biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.50.620 | HUPs | IPR014729 | Rossmann-like alpha/beta/alpha sandwich fold | 2 | 219 | 2.2E-74 |
| SUPERFAMILY | SSF52402 | Adenine nucleotide alpha hydrolases-like | - | - | 4 | 223 | 1.79E-64 |
| PANTHER | PTHR42914 | 7-CYANO-7-DEAZAGUANINE SYNTHASE | IPR018317 | Queuosine biosynthesis protein QueC | 1 | 223 | 1.3E-70 |
| Hamap | MF_01633 | 7-cyano-7-deazaguanine synthase [queC]. | IPR018317 | Queuosine biosynthesis protein QueC | 4 | 221 | 56.501068 |
| FunFam | G3DSA:3.40.50.620:FF:000131 | 7-cyano-7-deazaguanine synthase | - | - | 3 | 219 | 1.6E-124 |
| Pfam | PF06508 | Queuosine biosynthesis protein QueC | IPR018317 | Queuosine biosynthesis protein QueC | 5 | 215 | 5.3E-80 |
| CDD | cd01995 | ExsB | IPR018317 | Queuosine biosynthesis protein QueC | 5 | 214 | 7.8246E-85 |
| PIRSF | PIRSF006293 | ExsB | IPR018317 | Queuosine biosynthesis protein QueC | 2 | 224 | 1.6E-79 |
| NCBIfam | TIGR00364 | JCVI: 7-cyano-7-deazaguanine synthase QueC | IPR018317 | Queuosine biosynthesis protein QueC | 6 | 208 | 3.4E-62 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.