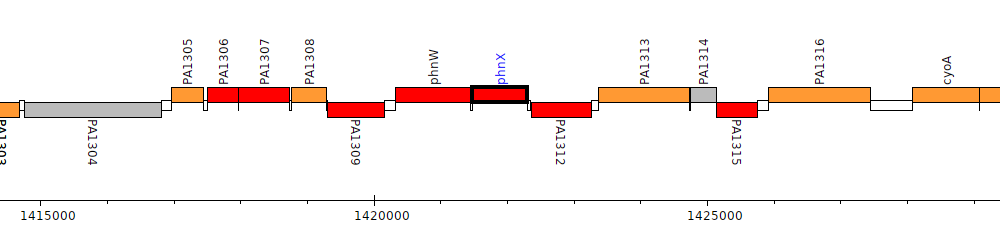
Pseudomonas aeruginosa PAO1, PA1311 (phnX)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0019700 | organic phosphonate catabolic process | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
9332393 | Reviewed by curator |
| Molecular Function | GO:0050194 | phosphonoacetaldehyde hydrolase activity | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
9332393 | Reviewed by curator |
| Molecular Function | GO:0050194 | phosphonoacetaldehyde hydrolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01375
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0019700 | organic phosphonate catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01375
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00440 | Phosphonate and phosphinate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | PHOSPHONOTASE-PWY | 2-aminoethylphosphonate degradation I | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| NCBIfam | TIGR01422 | JCVI: phosphonoacetaldehyde hydrolase | IPR006323 | Phosphonoacetaldehyde hydrolase | 9 | 259 | 3.3E-112 |
| FunFam | G3DSA:1.10.150.240:FF:000006 | Phosphonoacetaldehyde hydrolase | - | - | 27 | 104 | 5.3E-38 |
| SUPERFAMILY | SSF56784 | HAD-like | IPR036412 | HAD-like superfamily | 10 | 262 | 3.25E-57 |
| Gene3D | G3DSA:3.40.50.1000 | - | IPR023214 | HAD superfamily | 11 | 269 | 5.0E-96 |
| Gene3D | G3DSA:1.10.150.240 | Putative phosphatase; domain 2 | IPR023198 | Phosphoglycolate phosphatase-like, domain 2 | 27 | 104 | 5.0E-96 |
| PANTHER | PTHR43434 | PHOSPHOGLYCOLATE PHOSPHATASE | - | - | 10 | 257 | 4.1E-31 |
| SFLD | SFLDF00038 | phosphonoacetaldehyde hydrolase | - | - | 1 | 272 | 0.0 |
| Hamap | MF_01375 | Phosphonoacetaldehyde hydrolase [phnX]. | IPR006323 | Phosphonoacetaldehyde hydrolase | 1 | 275 | 142.689911 |
| SFLD | SFLDG01129 | C1.5: HAD, Beta-PGM, Phosphatase Like | - | - | 1 | 272 | 0.0 |
| NCBIfam | TIGR01509 | JCVI: HAD-IA family hydrolase | IPR006439 | HAD hydrolase, subfamily IA | 81 | 206 | 1.4E-10 |
| Pfam | PF00702 | haloacid dehalogenase-like hydrolase | - | - | 14 | 201 | 7.0E-7 |
| CDD | cd02586 | HAD_PHN | IPR006323 | Phosphonoacetaldehyde hydrolase | 9 | 250 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.