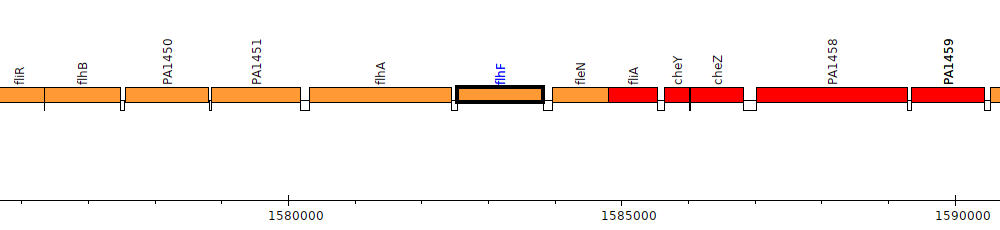
Pseudomonas aeruginosa PAO1, PA1453 (flhF)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0044781 | bacterial-type flagellum organization | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
17660418 | Reviewed by curator |
| Cellular Component | GO:0009288 | bacterial-type flagellum | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
1447978 | Reviewed by curator |
| Molecular Function | GO:0005525 | GTP binding | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
1447978 | Reviewed by curator |
| Biological Process | GO:0006614 | SRP-dependent cotranslational protein targeting to membrane |
Inferred from Sequence Model
Term mapped from: InterPro:SM00962
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016887 | ATPase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00382
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003924 | GTPase activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd17873
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005525 | GTP binding |
Inferred from Sequence Model
Term mapped from: InterPro:SM00962
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0044781 | bacterial-type flagellum organization |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03499
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Flagella assembly |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| NCBIfam | TIGR03499 | JCVI: flagellar biosynthesis protein FlhF | IPR020006 | Flagellar biosynthesis protein FlhF | 1 | 297 | 4.5E-76 |
| SUPERFAMILY | SSF52540 | P-loop containing nucleoside triphosphate hydrolases | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 207 | 398 | 2.64E-28 |
| Pfam | PF00448 | SRP54-type protein, GTPase domain | IPR000897 | Signal recognition particle, SRP54 subunit, GTPase domain | 211 | 399 | 5.1E-40 |
| Coils | Coil | Coil | - | - | 56 | 76 | - |
| SMART | SM00382 | AAA_5 | IPR003593 | AAA+ ATPase domain | 208 | 379 | 4.4E-6 |
| FunFam | G3DSA:3.40.50.300:FF:000695 | Flagellar biosynthesis regulator FlhF | - | - | 204 | 389 | 1.1E-61 |
| SMART | SM00962 | SRP54_3 | IPR000897 | Signal recognition particle, SRP54 subunit, GTPase domain | 209 | 401 | 2.9E-46 |
| Gene3D | G3DSA:1.20.120.1380 | Flagellar FlhF biosynthesis protein, N domain | - | - | 186 | 391 | 1.0E-44 |
| CDD | cd17873 | FlhF | IPR047040 | Flagellar biosynthesis protein FlhF, GTPase domain | 210 | 399 | 8.24826E-74 |
| PANTHER | PTHR43134 | SIGNAL RECOGNITION PARTICLE RECEPTOR SUBUNIT ALPHA | - | - | 188 | 398 | 1.6E-26 |
| Gene3D | G3DSA:3.40.50.300 | - | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 205 | 389 | 1.0E-44 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.