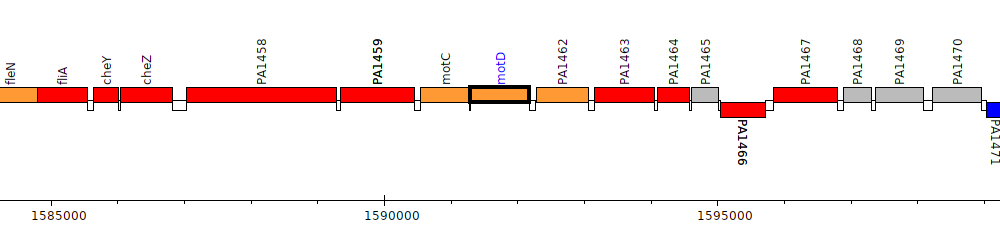
Pseudomonas aeruginosa PAO1, PA1461 (motD)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0071978 | bacterial-type flagellum-dependent swarming motility | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
15629949 | Reviewed by curator |
| Biological Process | GO:0071973 | bacterial-type flagellum-dependent cell motility | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
15375113 | Reviewed by curator |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Flagella assembly |
ECO:0000037
not_recorded |
|||
| KEGG | pae02030 | Bacterial chemotaxis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae02040 | Flagellar assembly | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Chemotaxis |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF103088 | OmpA-like | IPR036737 | OmpA-like domain superfamily | 127 | 240 | 1.7E-30 |
| CDD | cd07185 | OmpA_C-like | IPR006665 | OmpA-like domain | 134 | 240 | 1.11883E-33 |
| Pfam | PF00691 | OmpA family | IPR006665 | OmpA-like domain | 145 | 234 | 9.3E-17 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 249 | 296 | - |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 66 | 85 | - |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 63 | 96 | - |
| Gene3D | G3DSA:3.30.1330.60 | - | IPR036737 | OmpA-like domain superfamily | 77 | 255 | 1.2E-37 |
| Pfam | PF13677 | Membrane MotB of proton-channel complex MotA/MotB | IPR025713 | Motility protein B-like, N-terminal domain | 4 | 57 | 7.4E-22 |
| PANTHER | PTHR30329 | STATOR ELEMENT OF FLAGELLAR MOTOR COMPLEX | - | - | 2 | 242 | 2.9E-55 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.