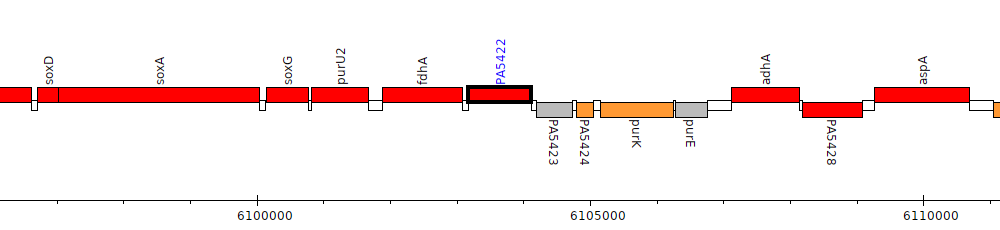
Pseudomonas aeruginosa PAO1, PA5422
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006798 | polyphosphate catabolic process | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
9734809 | Reviewed by curator |
| Molecular Function | GO:0008976 | polyphosphate kinase activity | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
9734809 | Reviewed by curator |
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF74650
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0005975 | carbohydrate metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:SSF74650
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0030246 | carbohydrate binding |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:2.70.98.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016853 | isomerase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01263
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00010 | Glycolysis / Gluconeogenesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd09020 | D-hex-6-P-epi_like | IPR025532 | Glucose-6-phosphate 1-epimerase | 33 | 310 | 1.70647E-118 |
| Pfam | PF01263 | Aldose 1-epimerase | IPR008183 | Aldose 1-/Glucose-6-phosphate 1-epimerase | 36 | 311 | 3.0E-62 |
| Gene3D | G3DSA:2.70.98.10 | - | IPR014718 | Glycoside hydrolase-type carbohydrate-binding | 9 | 311 | 1.1E-77 |
| PIRSF | PIRSF016020 | PHexose_mutarotase | IPR025532 | Glucose-6-phosphate 1-epimerase | 9 | 312 | 5.7E-50 |
| SUPERFAMILY | SSF74650 | Galactose mutarotase-like | IPR011013 | Galactose mutarotase-like domain superfamily | 21 | 309 | 9.42E-69 |
| PANTHER | PTHR11122 | APOSPORY-ASSOCIATED PROTEIN C-RELATED | - | - | 39 | 292 | 1.9E-48 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.