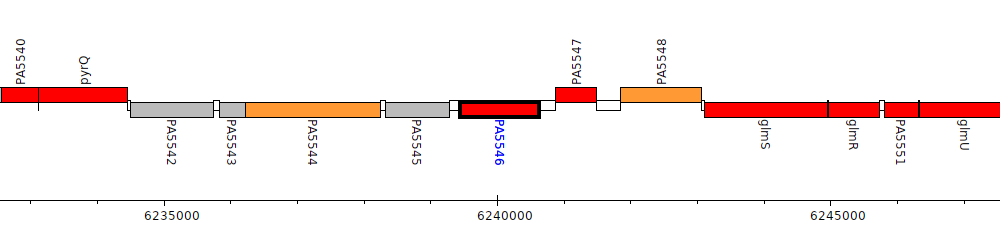
Pseudomonas aeruginosa PAO1, PA5546
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0008757 | S-adenosylmethionine-dependent methyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF53335
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0008610 | lipid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF003085
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM00828 | Methyltransferase in polyketide synthase (PKS) enzymes. | IPR020803 | Polyketide synthase, methyltransferase domain | 121 | 360 | 0.0077 |
| PIRSF | PIRSF003085 | CmaB | IPR003333 | Cyclopropane mycolic acid synthase | 1 | 394 | 1.2E-129 |
| Pfam | PF02353 | Mycolic acid cyclopropane synthetase | - | - | 98 | 372 | 2.9E-106 |
| NCBIfam | NF040703 | NCBIFAM: C17 cyclopropane fatty acid synthase CfaB | IPR048027 | Cyclopropane fatty acid synthase CfaB-like | 1 | 393 | 0.0 |
| SUPERFAMILY | SSF53335 | S-adenosyl-L-methionine-dependent methyltransferases | IPR029063 | S-adenosyl-L-methionine-dependent methyltransferase superfamily | 109 | 377 | 1.4E-85 |
| Gene3D | G3DSA:3.40.50.150 | Vaccinia Virus protein VP39 | IPR029063 | S-adenosyl-L-methionine-dependent methyltransferase superfamily | 92 | 375 | 1.2E-99 |
| PANTHER | PTHR43667 | CYCLOPROPANE-FATTY-ACYL-PHOSPHOLIPID SYNTHASE | - | - | 36 | 384 | 9.6E-103 |
| CDD | cd02440 | AdoMet_MTases | - | - | 162 | 262 | 2.53188E-15 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.