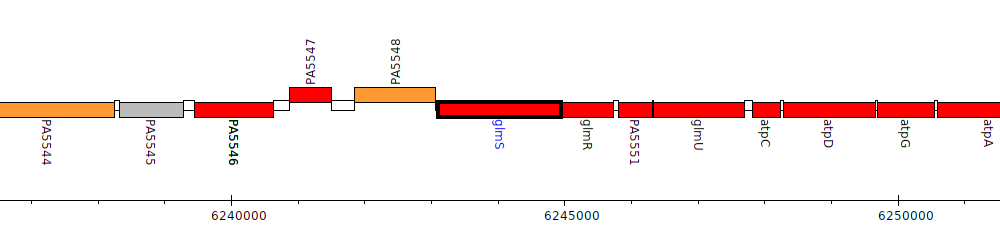
Pseudomonas aeruginosa PAO1, PA5549 (glmS)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004360 | glutamine-fructose-6-phosphate transaminase (isomerizing) activity | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
3297136 | Reviewed by curator |
| Biological Process | GO:0006048 | UDP-N-acetylglucosamine biosynthetic process | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
5541523 | Reviewed by curator |
| Biological Process | GO:1901135 | carbohydrate derivative metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:SSF53697
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004360 | glutamine-fructose-6-phosphate transaminase (isomerizing) activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00164
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:1901137 | carbohydrate derivative biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00164
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0097367 | carbohydrate derivative binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF53697
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCyc | UDPNAGSYN-PWY | UDP-N-acetyl-D-glucosamine biosynthesis I | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae00520 | Amino sugar and nucleotide sugar metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | PEP-LIPA-SYN-PWY | peptidoglycan and lipid A precursor biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Glutamate metabolism |
ECO:0000037
not_recorded |
|||
| PseudoCyc | OANTIGEN-PWY | O-antigen building blocks biosynthesis (E. coli) | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCAP | Aminosugars metabolism |
ECO:0000037
not_recorded |
|||
| PseudoCyc | KDO-PEP-LIPASYN-PWY | KDO<SUB>2</SUB>-lipid A and peptidoglycan biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae00250 | Alanine, aspartate and glutamate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd05008 | SIS_GlmS_GlmD_1 | IPR035466 | GlmS/AgaS, SIS domain 1 | 296 | 407 | 6.52998E-53 |
| FunFam | G3DSA:3.40.50.10490:FF:000001 | Glutamine--fructose-6-phosphate aminotransferase [isomerizing] | - | - | 248 | 450 | 2.5E-74 |
| Pfam | PF13522 | Glutamine amidotransferase domain | - | - | 67 | 175 | 8.5E-21 |
| NCBIfam | TIGR01135 | JCVI: glutamine--fructose-6-phosphate transaminase (isomerizing) | IPR005855 | Glucosamine-fructose-6-phosphate aminotransferase, isomerising | 2 | 611 | 0.0 |
| Pfam | PF01380 | SIS domain | IPR001347 | SIS domain | 464 | 593 | 1.7E-24 |
| Gene3D | G3DSA:3.40.50.10490 | - | - | - | 249 | 610 | 0.0 |
| CDD | cd05009 | SIS_GlmS_GlmD_2 | IPR035490 | GlmS/FrlB, SIS domain 2 | 457 | 609 | 1.98005E-68 |
| Hamap | MF_00164 | Glutamine--fructose-6-phosphate aminotransferase [isomerizing] [glmS]. | IPR005855 | Glucosamine-fructose-6-phosphate aminotransferase, isomerising | 1 | 611 | 40.724541 |
| Gene3D | G3DSA:3.40.50.10490 | - | - | - | 449 | 596 | 0.0 |
| CDD | cd00714 | GFAT | IPR047084 | Glucosamine-fructose-6-phosphate aminotransferase, isomerising, N-terminal domain | 2 | 216 | 2.22485E-125 |
| Pfam | PF01380 | SIS domain | IPR001347 | SIS domain | 291 | 418 | 1.2E-33 |
| SUPERFAMILY | SSF53697 | SIS domain | IPR046348 | SIS domain superfamily | 246 | 611 | 3.62E-118 |
| FunFam | G3DSA:3.60.20.10:FF:000006 | Glutamine--fructose-6-phosphate aminotransferase [isomerizing] | - | - | 2 | 240 | 1.1E-97 |
| FunFam | G3DSA:3.40.50.10490:FF:000002 | Glutamine--fructose-6-phosphate aminotransferase [isomerizing] | - | - | 451 | 596 | 3.9E-59 |
| Gene3D | G3DSA:3.60.20.10 | Glutamine Phosphoribosylpyrophosphate, subunit 1, domain 1 | IPR029055 | Nucleophile aminohydrolases, N-terminal | 2 | 241 | 1.4E-81 |
| PANTHER | PTHR10937 | GLUCOSAMINE--FRUCTOSE-6-PHOSPHATE AMINOTRANSFERASE, ISOMERIZING | - | - | 1 | 611 | 0.0 |
| SUPERFAMILY | SSF56235 | N-terminal nucleophile aminohydrolases (Ntn hydrolases) | IPR029055 | Nucleophile aminohydrolases, N-terminal | 2 | 235 | 2.14E-73 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.