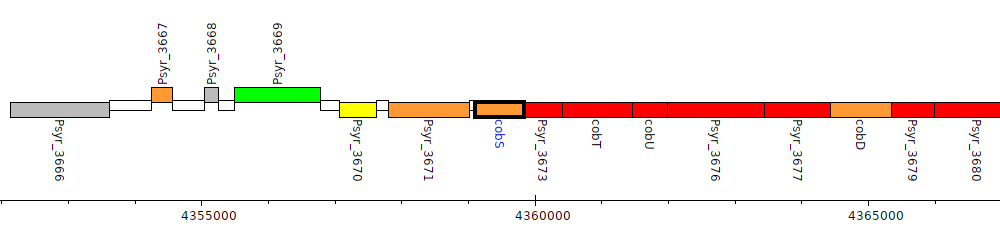
Pseudomonas syringae pv. syringae B728a, Psyr_3672 (cobS)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0051073 | adenosylcobinamide-GDP ribazoletransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF02654
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008818 | cobalamin 5'-phosphate synthase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF02654
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | psb00860 | Porphyrin and chlorophyll metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | superpathway of adenosylcobalamin salvage from cobinamide II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | 2-methyladeninyl adenosylcobamide biosynthesis from adenosylcobinamide-GDP | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | 5-methylbenzimidazolyl adenosylcobamide biosynthesis from adenosylcobinamide-GDP | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | 5-methoxybenzimidazolyl adenosylcobamide biosynthesis from adenosylcobinamide-GDP | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | adeninyl adenosylcobamide biosynthesis from adenosylcobinamide-GDP | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | benzimidazolyl adenosylcobamide biosynthesis from adenosylcobinamide-GDP | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | 5-methoxy-6-methylbenzimidazolyl adenosylcobamide biosynthesis from adenosylcobinamide-GDP | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | adenosylcobalamin biosynthesis from adenosylcobinamide-GDP II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | psb01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | 5-hydroxybenzimidazolyl adenosylcobamide biosynthesis from adenosylcobinamide-GDP | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | adenosylcobalamin biosynthesis from adenosylcobinamide-GDP I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | 4-methylphenyl adenosylcobamide biosynthesis from adenosylcobinamide-GDP | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | phenyl adenosylcobamide biosynthesis from adenosylcobinamide-GDP | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| NCBIfam | TIGR00317 | JCVI: adenosylcobinamide-GDP ribazoletransferase | IPR003805 | Adenosylcobinamide-GDP ribazoletransferase | 4 | 240 | 1.8E-35 |
| PANTHER | PTHR34148 | ADENOSYLCOBINAMIDE-GDP RIBAZOLETRANSFERASE | IPR003805 | Adenosylcobinamide-GDP ribazoletransferase | 4 | 242 | 2.8E-54 |
| Hamap | MF_00719 | Adenosylcobinamide-GDP ribazoletransferase [cobS]. | IPR003805 | Adenosylcobinamide-GDP ribazoletransferase | 1 | 243 | 48.497047 |
| Pfam | PF02654 | Cobalamin-5-phosphate synthase | IPR003805 | Adenosylcobinamide-GDP ribazoletransferase | 7 | 238 | 1.7E-50 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.