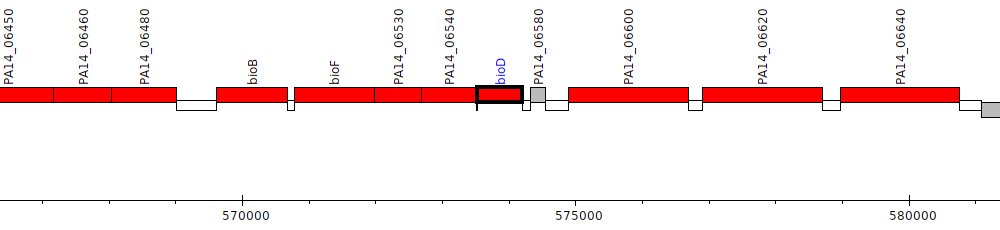
Pseudomonas aeruginosa UCBPP-PA14, PA14_06570 (bioD)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0000287 | magnesium ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00336
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004141 | dethiobiotin synthase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00336
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00336
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009102 | biotin biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00336
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | biotin biosynthesis from 8-amino-7-oxononanoate II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | pau00780 | Biotin metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR43210 | DETHIOBIOTIN SYNTHETASE | IPR004472 | Dethiobiotin synthase BioD | 1 | 217 | 1.0E-58 |
| Pfam | PF13500 | AAA domain | - | - | 4 | 212 | 3.1E-35 |
| Hamap | MF_00336 | ATP-dependent dethiobiotin synthetase BioD [bioD]. | IPR004472 | Dethiobiotin synthase BioD | 1 | 218 | 29.091362 |
| Gene3D | G3DSA:3.40.50.300 | - | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 3 | 222 | 6.5E-69 |
| FunFam | G3DSA:3.40.50.300:FF:000292 | ATP-dependent dethiobiotin synthetase BioD | - | - | 2 | 224 | 8.1E-80 |
| PIRSF | PIRSF006755 | DTB_synth | IPR004472 | Dethiobiotin synthase BioD | 1 | 226 | 2.1E-76 |
| SUPERFAMILY | SSF52540 | P-loop containing nucleoside triphosphate hydrolases | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 1 | 214 | 7.82E-60 |
| CDD | cd03109 | DTBS | - | - | 2 | 194 | 4.2344E-47 |
| NCBIfam | TIGR00347 | JCVI: dethiobiotin synthase | IPR004472 | Dethiobiotin synthase BioD | 5 | 176 | 2.9E-44 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.