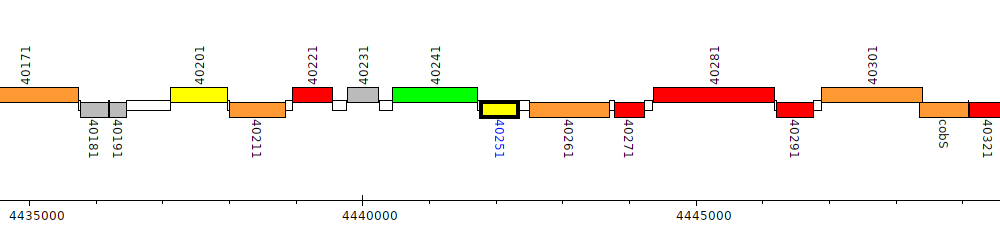
Pseudomonas aeruginosa LESB58, PALES_40251
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004602 | glutathione peroxidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR01011
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006979 | response to oxidative stress |
Inferred from Sequence Model
Term mapped from: InterPro:PR01011
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd00340 | GSH_Peroxidase | IPR000889 | Glutathione peroxidase | 37 | 180 | 3.85183E-68 |
| FunFam | G3DSA:3.40.30.10:FF:000246 | Glutathione peroxidase | - | - | 18 | 183 | 1.9E-94 |
| PIRSF | PIRSF000303 | Glutathion_perox | IPR000889 | Glutathione peroxidase | 1 | 184 | 3.7E-56 |
| PANTHER | PTHR11592 | GLUTATHIONE PEROXIDASE | IPR000889 | Glutathione peroxidase | 40 | 179 | 9.9E-31 |
| Pfam | PF00255 | Glutathione peroxidase | IPR000889 | Glutathione peroxidase | 39 | 134 | 1.8E-22 |
| SUPERFAMILY | SSF52833 | Thioredoxin-like | IPR036249 | Thioredoxin-like superfamily | 34 | 180 | 2.28E-42 |
| PRINTS | PR01011 | Glutathione peroxidase family signature | IPR000889 | Glutathione peroxidase | 82 | 98 | 2.9E-9 |
| PRINTS | PR01011 | Glutathione peroxidase family signature | IPR000889 | Glutathione peroxidase | 47 | 64 | 2.9E-9 |
| Gene3D | G3DSA:3.40.30.10 | Glutaredoxin | - | - | 23 | 183 | 7.0E-43 |
| PRINTS | PR01011 | Glutathione peroxidase family signature | IPR000889 | Glutathione peroxidase | 142 | 151 | 2.9E-9 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.