Pseudomonas aeruginosa PAO1, PA0170 (siaC)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
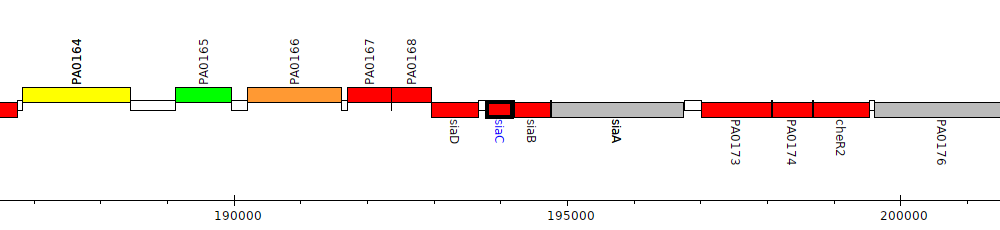
Gene Feature Overview
| Strain |
Pseudomonas aeruginosa PAO1 (Stover et al., 2000)
GCF_000006765.1|latest |
| Locus Tag |
PA0170
|
| Name |
siaC
|
| Replicon | chromosome |
| Genomic location | 193799 - 194179 (- strand) |
| Transposon Mutants | 2 transposon mutants in PAO1 |
| Comment | The SiaC protein shares structural similarities with anti-sigma factor antagonists (PDB-ID: 6K4F)[PMID:33156827] |
| Comment | The phosphorylation status of T68 of the SiaC protein is balanced by the phosphatase activity of SiaA (PA0172) and kinase activity of SiaB (PA0171)[PMID:33156827] |
| Comment | The SiaABC threonine phosphorylation pathway controls aggregate/biofilm formation in response to carbon availability (glucose, succinate, ethanol, 2,3-butanediol)[PMID:33156827] |
Cross-References
| RefSeq | NP_248860.1 |
| GI | 15595368 |
| Affymetrix | PA0170_at |
| DNASU | PaCD00006075 |
| Entrez | 878493 |
| GenBank | AAG03560.1 |
| INSDC | AAG03560.1 |
| NCBI Locus Tag | PA0170 |
| protein_id(GenBank) | gb|AAG03560.1|AE004455_1|gnl|PseudoCAP|PA0170 |
| TIGR | NTL03PA00171 |
| UniParc | UPI00000C4F79 |
| UniProtKB Acc | Q9I6W2 |
| UniProtKB ID | Q9I6W2_PSEAE |
| UniRef100 | UniRef100_Q02UQ8 |
| UniRef50 | UniRef50_Q02UQ8 |
| UniRef90 | UniRef90_Q02UQ8 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
SiaC
|
| Synonyms |
putative anti-sigma factor antagonist |
| Evidence for Translation |
Identified using nanoflow high-pressure liquid chromatography (HPLC) in conjunction with microelectrospray ionization on LTQ XL mass spectrometer (PMID:24291602).
|
| Charge (pH 7) | -11.07 |
| Kyte-Doolittle Hydrophobicity Value | -0.552 |
| Molecular Weight (kDa) | 14.6 |
| Isoelectric Point (pI) | 4.29 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
PDB 3D Structures
| Accession | Header | Accession Date | Compound | Source | Resolution | Method | Percent Identity |
| 7E6G | DE NOVO PROTEIN | 02/22/21 | Crystal structure of diguanylate cyclase SiaD in complex with its activator SiaC from Pseudomonas aeruginosa | Pseudomonas aeruginosa | 2.65 | X-RAY DIFFRACTION | 100.0 |
| 6KKO | GENE REGULATION | 07/26/19 | The crystal structure of SiaB-SiaC complex from Pseudomonas aeruginosa | Pseudomonas aeruginosa | 2.099 | X-RAY DIFFRACTION | 100.0 |
| 6KKP | GENE REGULATION | 07/26/19 | The crystal structure of apo-SiaC from Pseudomonas aeruginosa | Pseudomonas aeruginosa | 2.5 | X-RAY DIFFRACTION | 100.0 |
| 6K4F | BIOSYNTHETIC PROTEIN | 05/23/19 | SiaC of Pseudomonas aeruginosa | Pseudomonas aeruginosa | 1.735 | X-RAY DIFFRACTION | 100.0 |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 53 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG000168 (328 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PA0170
Search term: siaC
Search term: SiaC
|
Human Homologs
References
|
The SiaABC threonine phosphorylation pathway controls biofilm formation in response to carbon availability in Pseudomonas aeruginosa.
Poh WH, Lin J, Colley B, Müller N, Goh BC, Schleheck D, El Sahili A, Marquardt A, Liang Y, Kjelleberg S, Lescar J, Rice SA, Klebensberger J
PLoS One 2020;15(11):e0241019
PubMed ID: 33156827
|