Pseudomonas aeruginosa UCBPP-PA14, PA14_06750 (nirS)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
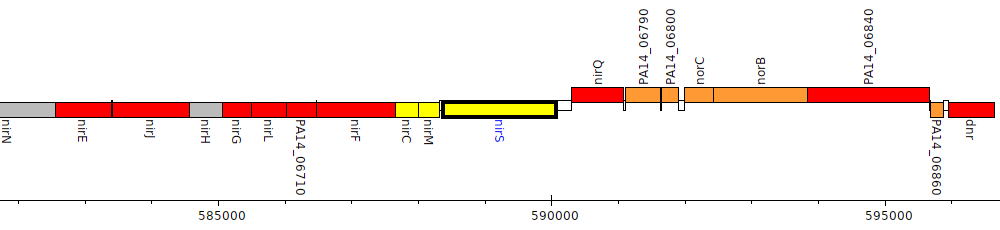
Gene Feature Overview
| Strain |
Pseudomonas aeruginosa UCBPP-PA14 (Lee et al., 2006)
GCF_000014625.1|latest |
| Locus Tag |
PA14_06750
|
| Name |
nirS
|
| Replicon | chromosome |
| Genomic location | 588371 - 590077 (- strand) |
| Transposon Mutants | 2 transposon mutants in UCBPP-PA14 |
| Transposon Mutants in orthologs | 13 transposon mutants in orthologs |
Cross-References
| RefSeq | YP_788689.1 |
| GI | 116054245 |
| Entrez | 4383885 |
| INSDC | ABJ15481.1 |
| NCBI Locus Tag | PA14_06750 |
| UniParc | UPI000045D429 |
| UniProtKB Acc | Q02TP7 |
| UniProtKB ID | Q02TP7_PSEAB |
| UniRef100 | UniRef100_Q02TP7 |
| UniRef50 | UniRef50_P24474 |
| UniRef90 | UniRef90_P24474 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
nitrite reductase
|
| Synonyms | |
| Evidence for Translation | |
| Charge (pH 7) | 5.04 |
| Kyte-Doolittle Hydrophobicity Value | -0.395 |
| Molecular Weight (kDa) | 62.7 |
| Isoelectric Point (pI) | 8.21 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
PDB 3D Structures
| Accession | Header | Accession Date | Compound | Source | Resolution | Method | Percent Identity |
| 5GUW | MEMBRANE PROTEIN | 08/31/16 | Complex of Cytochrome cd1 Nitrite Reductase and Nitric Oxide Reductase in Denitrification of Pseudomonas aeruginosa | Pseudomonas aeruginosa PAO1 | 3.2 | X-RAY DIFFRACTION | 99.6 |
| 1N15 | OXIDOREDUCTASE | 09/04/98 | FOLLOWING THE C HEME REDUCTION IN NITRITE REDUCTASE FROM PSEUDOMONAS AERUGINOSA | Pseudomonas aeruginosa | 2.9 | X-RAY DIFFRACTION | 99.6 |
| 1N90 | OXIDOREDUCTASE | 09/04/98 | FOLLOWING THE C HEME REDUCTION IN NITRITE REDUCTASE FROM PSEUDOMONAS AERUGINOSA | Pseudomonas aeruginosa | 2.9 | X-RAY DIFFRACTION | 99.6 |
| 1HZU | OXIDOREDUCTASE | 01/26/01 | DOMAIN SWING UPON HIS TO ALA MUTATION IN NITRITE REDUCTASE OF PSEUDOMONAS AERUGINOSA | Pseudomonas aeruginosa | 2.7 | X-RAY DIFFRACTION | 99.4 |
| 1GJQ | OXIDOREDUCTASE | 08/01/01 | Pseudomonas aeruginosa cd1 nitrite reductase reduced cyanide complex | PSEUDOMONAS AERUGINOSA | 2.7 | X-RAY DIFFRACTION | 99.6 |
| 6TPO | OXIDOREDUCTASE | 12/13/19 | Conformation of cd1 nitrite reductase NirS without bound heme d1 | Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) | 1.861 | X-RAY DIFFRACTION | 99.6 |
| 1BL9 | OXIDOREDUCTASE | 07/20/98 | CONFORMATIONAL CHANGES OCCURRING UPON REDUCTION IN NITRITE REDUCTASE FROM PSEUDOMONAS AERUGINOSA | Pseudomonas aeruginosa | 2.9 | X-RAY DIFFRACTION | 99.6 |
| 1NNO | OXIDOREDUCTASE | 07/20/98 | CONFORMATIONAL CHANGES OCCURRING UPON NO BINDING IN NITRITE REDUCTASE FROM PSEUDOMONAS AERUGINOSA | Pseudomonas aeruginosa | 2.65 | X-RAY DIFFRACTION | 99.6 |
| 1N50 | OXIDOREDUCTASE | 09/04/98 | FOLLOWING THE C HEME REDUCTION IN NITRITE REDUCTASE FROM PSEUDOMONAS AERUGINOSA | Pseudomonas aeruginosa | 2.9 | X-RAY DIFFRACTION | 99.6 |
| 6TSI | OXIDOREDUCTASE | 12/20/19 | cd1 nitrite reductase NirS with bound dihydro-heme d1 | Pseudomonas aeruginosa PAO1 | 2.38 | X-RAY DIFFRACTION | 99.6 |
| 1NIR | NITRITE REDUCTASE | 06/17/97 | OXYDIZED NITRITE REDUCTASE FROM PSEUDOMONAS AERUGINOSA | Pseudomonas aeruginosa | 2.15 | X-RAY DIFFRACTION | 99.6 |
| 1HZV | OXIDOREDUCTASE | 01/26/01 | DOMAIN SWING UPON HIS TO ALA MUTATION IN NITRITE REDUCTASE OF PSEUDOMONAS AERUGINOSA | Pseudomonas aeruginosa | 2.83 | X-RAY DIFFRACTION | 99.4 |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 48 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG000503 (354 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PA14_06750
Search term: nirS
Search term: nitrite reductase
|
Human Homologs
References
No references are associated with this feature.