Pseudomonas aeruginosa PAO1, PA0845 (cerN)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
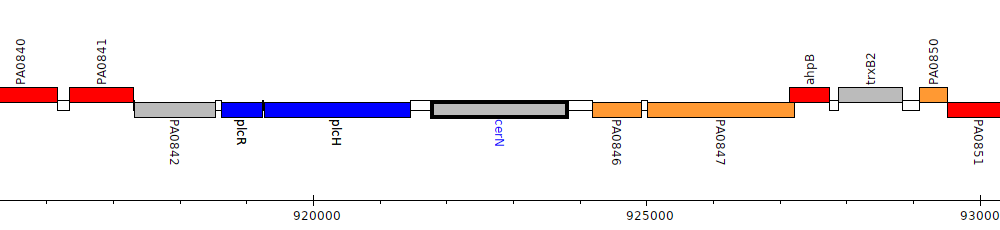
Gene Feature Overview
| Strain |
Pseudomonas aeruginosa PAO1 (Stover et al., 2000)
GCF_000006765.1|latest |
| Locus Tag |
PA0845
|
| Name |
cerN
|
| Replicon | chromosome |
| Genomic location | 921792 - 923804 (- strand) |
| Transposon Mutants | 3 transposon mutants in PAO1 |
| Transposon Mutants in orthologs | 2 transposon mutants in orthologs |
| Comment | Neutral ceramidase regulated by the sphingosine-sensing transcription factor SphR |
Cross-References
| RefSeq | NP_249536.1 |
| GI | 15596042 |
| Affymetrix | PA0845_at |
| Entrez | 880698 |
| GenBank | AAG04234.1 |
| NCBI Locus Tag | PA0845 |
| protein_id(GenBank) | gb|AAG04234.1|AE004519_5|gnl|PseudoCAP|PA0845 |
| TIGR | NTL03PA00846 |
| UniProtKB Acc | Q9I596 |
| UniProtKB ID | NCASE_PSEAE |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
CerN
|
| Synonyms | |
| Evidence for Translation |
Identified using nanoflow high-pressure liquid chromatography (HPLC) in conjunction with microelectrospray ionization on LTQ XL mass spectrometer (PMID:24291602).
|
| Charge (pH 7) | -6.76 |
| Kyte-Doolittle Hydrophobicity Value | -0.266 |
| Molecular Weight (kDa) | 73.4 |
| Isoelectric Point (pI) | 6.17 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
PDB 3D Structures
| Accession | Header | Accession Date | Compound | Source | Resolution | Method | Percent Identity |
| 2ZWS | HYDROLASE | 12/17/08 | Crystal Structure Analysis of neutral ceramidase from Pseudomonas aeruginosa | Pseudomonas aeruginosa | 1.4 | X-RAY DIFFRACTION | 99.5 |
| 2ZXC | HYDROLASE | 12/22/08 | Ceramidase complexed with C2 | Pseudomonas aeruginosa | 2.2 | X-RAY DIFFRACTION | 99.5 |
Drugs Targeting this Protein
Identified by Diamond using e-value cutoff of 0.0001 and returning alignments that span 100% of the query sequence and that have more than 95% identity.
| Drug Name | Source Accession | Source DB | Version | Target Accession | Target Description | Percent Identity | Alignment Length | E-Value |
| N-[(1R,2R,3E)-2-hydroxy-1-(hydroxymethyl)heptadec-3-en-1-yl]acetamide | DB06960 | DrugBank | 5.1.4 | Q9I596 | Neutral ceramidase | 100.0 | 670 | 0.0e+00 |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 41 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG000812 (298 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PA0845
Search term: cerN
Search term: CerN
|
Human Homologs
|
Ensembl
110, assembly
GRCh38.p14
|
N-acylsphingosine amidohydrolase 2 [Source:HGNC Symbol;Acc:HGNC:18860]
E-value:
1.2e-117
Percent Identity:
36.1
|
|
Ensembl
110, assembly
GRCh38.p14
|
N-acylsphingosine amidohydrolase 2 [Source:HGNC Symbol;Acc:HGNC:18860]
E-value:
1.2e-117
Percent Identity:
36.1
|
References
|
Genome-wide identification of Pseudomonas aeruginosa exported proteins using a consensus computational strategy combined with a laboratory-based PhoA fusion screen.
Lewenza S, Gardy JL, Brinkman FS, Hancock RE
Genome Res 2005 Feb;15(2):321-9
PubMed ID: 15687295
|
|
Detection of host-derived sphingosine by Pseudomonas aeruginosa is important for survival in the murine lung.
LaBauve AE, Wargo MJ
PLoS Pathog 2014 Jan;10(1):e1003889
PubMed ID: 24465209
|
|
Molecular cloning, sequencing, and expression of the gene encoding alkaline ceramidase from Pseudomonas aeruginosa. Cloning of a ceramidase homologue from Mycobacterium tuberculosis.
Okino N, Ichinose S, Omori A, Imayama S, Nakamura T, Ito M
J Biol Chem 1999 Dec 17;274(51):36616-22
PubMed ID: 10593963
|
|
Purification and characterization of a novel ceramidase from Pseudomonas aeruginosa.
Okino N, Tani M, Imayama S, Ito M
J Biol Chem 1998 Jun 5;273(23):14368-73
PubMed ID: 9603946
|