Pseudomonas fluorescens F113, PSF113_5352
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
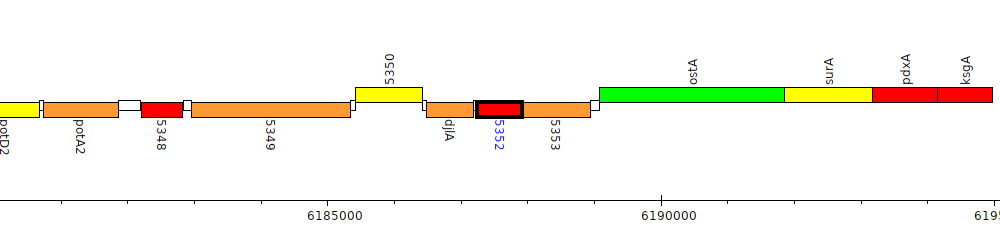
Gene Feature Overview
| Strain |
Pseudomonas fluorescens F113
GCF_000237065.1|latest |
| Locus Tag |
PSF113_5352
|
| Name |
|
| Replicon | chromosome |
| Genomic location | 6187246 - 6187917 (- strand) |
Cross-References
| RefSeq | YP_005210719.1 |
| GI | 378953231 |
| EC Number | 2.7.7.- |
| Entrez | 11833467 |
| NCBI Locus Tag | PSF113_5352 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
nucleotidyltransferase
|
| Synonyms | |
| Evidence for Translation | |
| Charge (pH 7) | -6.14 |
| Kyte-Doolittle Hydrophobicity Value | -0.058 |
| Molecular Weight (kDa) | 24162.4 |
| Isoelectric Point (pI) | 6.07 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
AlphaFold 2 Protein Structure Predictions
Protein structure predictions using a neural network model developed by DeepMind. If a UniProtKB accession is associated with this protein, a search link will be provided below.
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 611 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG000580 (536 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PSF113_5352
Search term: nucleotidyltransferase
|
Human Homologs
|
Ensembl
110, assembly
GRCh38.p14
|
GDP-mannose pyrophosphorylase B [Source:HGNC Symbol;Acc:HGNC:22932]
E-value:
1.8e-22
Percent Identity:
31.8
|
|
Ensembl
110, assembly
GRCh38.p14
|
GDP-mannose pyrophosphorylase B [Source:HGNC Symbol;Acc:HGNC:22932]
E-value:
1.8e-22
Percent Identity:
31.8
|
|
Ensembl
110, assembly
GRCh38.p14
|
GDP-mannose pyrophosphorylase B [Source:HGNC Symbol;Acc:HGNC:22932]
E-value:
1.8e-22
Percent Identity:
31.8
|
References
No references are associated with this feature.