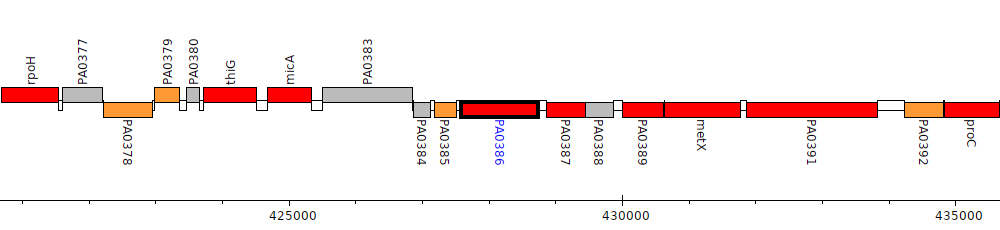
Pseudomonas aeruginosa PAO1, PA0386
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0051536 | iron-sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:SM00729
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00729
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006779 | porphyrin-containing compound biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:SFLDF00562
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004109 | coproporphyrinogen oxidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SFLDF00562
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051539 | 4 iron, 4 sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:SFLDF00562
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005737 | cytoplasm |
Inferred from Sequence Model
Term mapped from: InterPro:SFLDF00562
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SFLD | SFLDG01082 | B12-binding domain containing | - | - | 21 | 252 | 2.5E-17 |
| Pfam | PF04055 | Radical SAM superfamily | IPR007197 | Radical SAM | 16 | 183 | 1.2E-19 |
| Gene3D | G3DSA:3.20.20.70 | Aldolase class I | IPR013785 | Aldolase-type TIM barrel | 14 | 223 | 1.2E-10 |
| PANTHER | PTHR13932 | COPROPORPHYRINIGEN III OXIDASE | IPR034505 | Anaerobic coproporphyrinogen-III oxidase | 10 | 383 | 6.0E-111 |
| CDD | cd01335 | Radical_SAM | - | - | 20 | 222 | 3.58228E-14 |
| Pfam | PF06969 | HemN C-terminal domain | IPR010723 | HemN, C-terminal | 309 | 373 | 5.8E-14 |
| NCBIfam | TIGR00539 | JCVI: radical SAM family heme chaperone HemW | IPR004559 | Heme chaperone HemW-like | 13 | 346 | 1.2E-117 |
| SFLD | SFLDF00288 | HemN-like, clustered with nucleoside-triphosphate RdgB | - | - | 7 | 383 | 0.0 |
| SFLD | SFLDF00562 | HemN-like, clustered with heat shock genes | IPR004559 | Heme chaperone HemW-like | 11 | 383 | 0.0 |
| SUPERFAMILY | SSF102114 | Radical SAM enzymes | - | - | 10 | 383 | 5.49E-107 |
| SFLD | SFLDG01065 | anaerobic coproporphyrinogen-III oxidase like | - | - | 7 | 383 | 0.0 |
| SMART | SM00729 | MiaB | IPR006638 | Elp3/MiaA/NifB-like, radical SAM core domain | 11 | 227 | 4.3E-57 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.