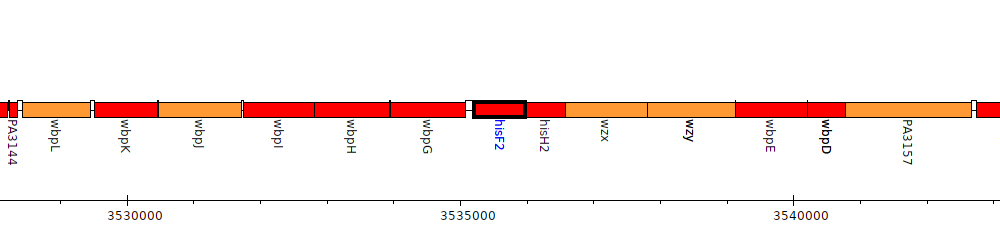
Pseudomonas aeruginosa PAO1, PA3151 (hisF2)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006520 | cellular amino acid metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006547 | histidine metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0009243 | O antigen biosynthetic process | Inferred from Experiment | ECO:0000269 experimental evidence used in manual assertion |
18621892 | Reviewed by curator |
| Biological Process | GO:0000105 | histidine biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:cd04731
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000107 | imidazoleglycerol-phosphate synthase activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd04731
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016833 | oxo-acid-lyase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03572
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01230 | Biosynthesis of amino acids | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | PRPP-PWY | superpathway of histidine, purine, and pyrimidine biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00340 | Histidine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | HISTSYN-PWY | L-histidine biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Histidine metabolism |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| NCBIfam | NF038364 | NCBIFAM: AglZ/HisF2 family acetamidino modification protein | - | - | 3 | 248 | 1.0E-113 |
| SUPERFAMILY | SSF51366 | Ribulose-phoshate binding barrel | IPR011060 | Ribulose-phosphate binding barrel | 1 | 232 | 1.02E-61 |
| Pfam | PF00977 | Histidine biosynthesis protein | IPR006062 | Histidine biosynthesis protein | 5 | 232 | 3.7E-66 |
| Gene3D | G3DSA:3.20.20.70 | Aldolase class I | IPR013785 | Aldolase-type TIM barrel | 1 | 243 | 2.0E-81 |
| NCBIfam | TIGR03572 | JCVI: glycosyl amidation-associated protein WbuZ | IPR020021 | Glycosyl amidation-associated protein, WbuZ | 1 | 231 | 2.0E-98 |
| CDD | cd04731 | HisF | IPR004651 | Histidine biosynthesis, HisF | 4 | 243 | 1.1274E-126 |
| PANTHER | PTHR21235 | IMIDAZOLE GLYCEROL PHOSPHATE SYNTHASE SUBUNIT HISF/H IGP SYNTHASE SUBUNIT HISF/H | - | - | 2 | 230 | 4.8E-61 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.