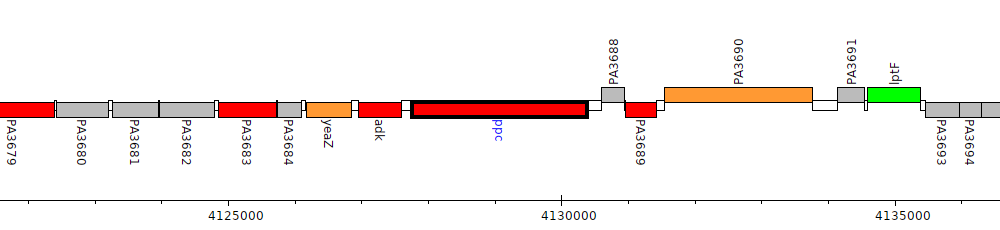
Pseudomonas aeruginosa PAO1, PA3687 (ppc)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006091 | generation of precursor metabolites and energy | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0015977 | carbon fixation | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0008152 | metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0016814 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in cyclic amidines | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006090 | pyruvate metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0019643 | reductive tricarboxylic acid cycle | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0008964 | phosphoenolpyruvate carboxylase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR00150
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF51621
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0015977 | carbon fixation |
Inferred from Sequence Model
Term mapped from: InterPro:PR00150
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006099 | tricarboxylic acid cycle |
Inferred from Sequence Model
Term mapped from: InterPro:PR00150
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01200 | Carbon metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | formaldehyde assimilation I (serine pathway) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | pae00620 | Pyruvate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | C4 photosynthetic carbon assimilation cycle, NADP-ME type | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | C4 photosynthetic carbon assimilation cycle, NAD-ME type | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | P22-PWY | acetyl-CoA assimilation | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae00680 | Methane metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | FERMENTATION-PWY | mixed acid fermentation | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| MetaCyc | L-glutamine biosynthesis III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | gluconeogenesis II (<i>Methanobacterium thermoautotrophicum</i>) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | ethylene biosynthesis V (engineered) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| PseudoCAP | Reductive carboxylate cycle (CO2 fixation) |
ECO:0000037
not_recorded |
|||
| MetaCyc | <i>Methanobacterium thermoautotrophicum</i> biosynthetic metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| PseudoCAP | Pyruvate metabolism |
ECO:0000037
not_recorded |
|||
| PseudoCyc | P106-PWY | serine-isocitrate lyase pathway | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | ANARESP1-PWY | anaerobic respiration | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| MetaCyc | partial TCA cycle (obligate autotrophs) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| PseudoCAP | Carbon fixation |
ECO:0000037
not_recorded |
|||
| MetaCyc | C4 photosynthetic carbon assimilation cycle, PEPCK type | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00150 | Phosphoenolpyruvate carboxylase signature | IPR021135 | Phosphoenolpyruvate carboxylase | 135 | 148 | 8.9E-69 |
| Hamap | MF_00595 | Phosphoenolpyruvate carboxylase [ppc]. | IPR022805 | Phosphoenolpyruvate carboxylase, bacterial/plant-type | 1 | 878 | 18.767492 |
| PRINTS | PR00150 | Phosphoenolpyruvate carboxylase signature | IPR021135 | Phosphoenolpyruvate carboxylase | 385 | 405 | 8.9E-69 |
| SUPERFAMILY | SSF51621 | Phosphoenolpyruvate/pyruvate domain | IPR015813 | Pyruvate/Phosphoenolpyruvate kinase-like domain superfamily | 5 | 878 | 0.0 |
| Pfam | PF00311 | Phosphoenolpyruvate carboxylase | IPR021135 | Phosphoenolpyruvate carboxylase | 8 | 878 | 0.0 |
| Coils | Coil | Coil | - | - | 762 | 782 | - |
| PRINTS | PR00150 | Phosphoenolpyruvate carboxylase signature | IPR021135 | Phosphoenolpyruvate carboxylase | 247 | 262 | 8.9E-69 |
| PRINTS | PR00150 | Phosphoenolpyruvate carboxylase signature | IPR021135 | Phosphoenolpyruvate carboxylase | 534 | 554 | 8.9E-69 |
| Gene3D | G3DSA:1.20.1440.90 | Phosphoenolpyruvate/pyruvate domain | - | - | 275 | 387 | 4.4E-28 |
| PRINTS | PR00150 | Phosphoenolpyruvate carboxylase signature | IPR021135 | Phosphoenolpyruvate carboxylase | 576 | 605 | 8.9E-69 |
| PANTHER | PTHR30523 | PHOSPHOENOLPYRUVATE CARBOXYLASE | IPR021135 | Phosphoenolpyruvate carboxylase | 5 | 878 | 0.0 |
| PRINTS | PR00150 | Phosphoenolpyruvate carboxylase signature | IPR021135 | Phosphoenolpyruvate carboxylase | 188 | 204 | 8.9E-69 |
| PRINTS | PR00150 | Phosphoenolpyruvate carboxylase signature | IPR021135 | Phosphoenolpyruvate carboxylase | 707 | 733 | 8.9E-69 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.