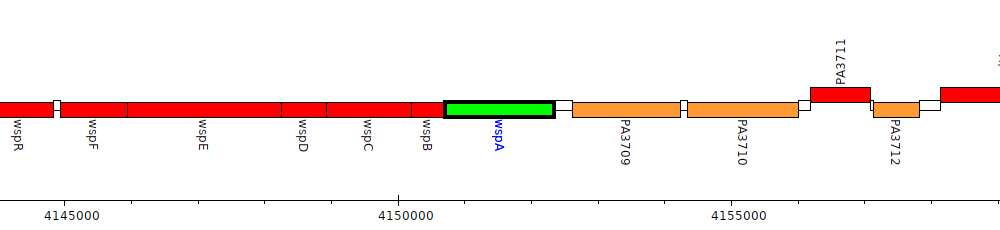
Pseudomonas aeruginosa PAO1, PA3708 (wspA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006935 | chemotaxis |
Inferred from Sequence Model
Term mapped from: InterPro:PR00260
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004888 | transmembrane signaling receptor activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR00260
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0007165 | signal transduction |
Inferred from Sequence Model
Term mapped from: InterPro:SM00283
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:SM00283
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Chemotactic transducer (MCP) |
ECO:0000037
not_recorded |
|||
| KEGG | pae02025 | Biofilm formation - Pseudomonas aeruginosa | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae02020 | Two-component system | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Chemosensory |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00260 | Bacterial chemotaxis sensory transducer signature | IPR004090 | Chemotaxis methyl-accepting receptor | 378 | 405 | 4.9E-17 |
| PRINTS | PR00260 | Bacterial chemotaxis sensory transducer signature | IPR004090 | Chemotaxis methyl-accepting receptor | 407 | 436 | 4.9E-17 |
| Pfam | PF00672 | HAMP domain | IPR003660 | HAMP domain | 210 | 261 | 1.2E-10 |
| SMART | SM00304 | HAMP_11 | IPR003660 | HAMP domain | 213 | 265 | 1.4E-11 |
| SUPERFAMILY | SSF58104 | Methyl-accepting chemotaxis protein (MCP) signaling domain | - | - | 233 | 542 | 1.83E-61 |
| Pfam | PF00015 | Methyl-accepting chemotaxis protein (MCP) signalling domain | IPR004089 | Methyl-accepting chemotaxis protein (MCP) signalling domain | 348 | 505 | 5.7E-34 |
| CDD | cd06225 | HAMP | - | - | 216 | 260 | 7.63391E-10 |
| CDD | cd19411 | MCP2201-like_sensor | IPR047347 | Putative sensory transducer protein YvaQ-like, ligand-binding sensor domain | 73 | 182 | 4.58101E-9 |
| PANTHER | PTHR32089 | METHYL-ACCEPTING CHEMOTAXIS PROTEIN MCPB | - | - | 10 | 541 | 1.6E-82 |
| PRINTS | PR00260 | Bacterial chemotaxis sensory transducer signature | IPR004090 | Chemotaxis methyl-accepting receptor | 301 | 330 | 4.9E-17 |
| Gene3D | G3DSA:1.10.287.950 | - | - | - | 213 | 542 | 1.5E-69 |
| SMART | SM00283 | MA_2 | IPR004089 | Methyl-accepting chemotaxis protein (MCP) signalling domain | 273 | 541 | 2.4E-46 |
| Pfam | PF12729 | Four helix bundle sensory module for signal transduction | IPR024478 | Chemotaxis methyl-accepting receptor HlyB-like, 4HB MCP domain | 3 | 182 | 1.1E-21 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.